3.1.1 Soil as a Living System and the Foundation of Planetary Health
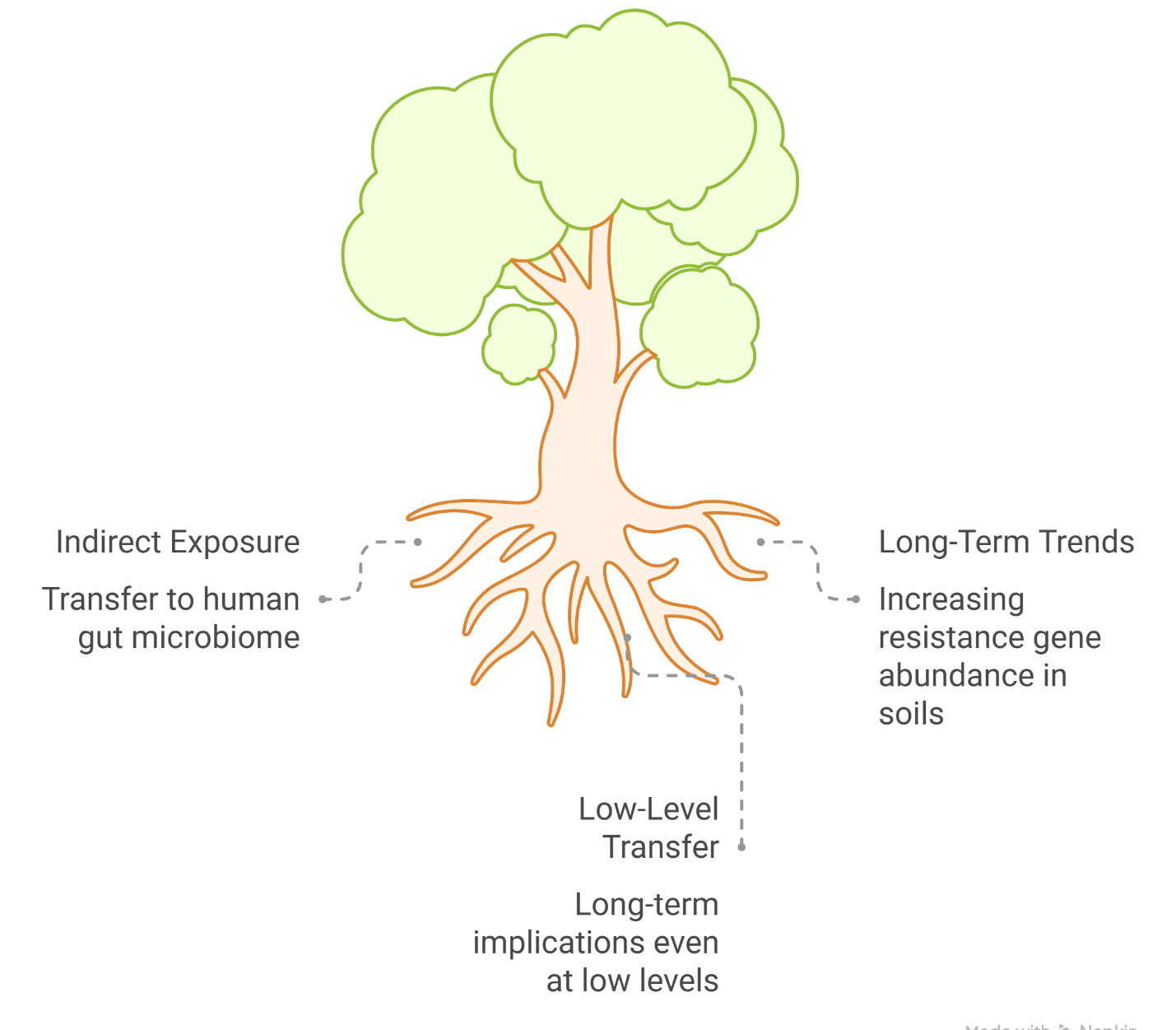
Soil, despite often being treated as a passive substrate in agricultural discourse, is anything but inert. It is, in many respects, a living system—densely populated, metabolically active, and astonishingly complex. Within a single gram, one may find billions of microbial cells and an almost incomprehensible diversity of taxa, forming what is sometimes described as an “unseen majority.” These microbial consortia are not merely present; they actively regulate biogeochemical cycles, including carbon sequestration and nitrogen transformation, processes that underpin the stability of terrestrial ecosystems. It is within this broader ecological framing that the concept of Planetary Health becomes particularly relevant, emphasizing that human well-being is deeply—and perhaps unavoidably—intertwined with the integrity of natural systems (Zhu et al., 2019; EFSA Panel on Biological Hazards, 2021). Yet, this intricate biological network is increasingly subjected to anthropogenic pressures. The growing reliance on reclaimed wastewater and organic amendments such as manure introduces not only nutrients but also a suite of emerging contaminants, including antibiotics and antibiotic resistance determinants. Over time, these inputs may subtly—but persistently—reshape the structure and function of soil microbial communities, raising concerns about long-term ecological stability (Phan et al., 2024; Wu et al., 2025). Figure 1 depicting the environmental dissemination of antimicrobial resistance across the soil–plant–water continuum in reclaimed wastewater–irrigated rice ecosystems, highlighting microbial, ecological, and human health linkages.
3.1.2 From Intrinsic Resistome to Anthropogenic Amplification
It is important, however, to distinguish between what might be termed the intrinsic soil resistome and the more recent, human-driven amplification of resistance. Soil microorganisms have, for millions of years, evolved mechanisms to resist naturally occurring antibiotics, using them as tools in ecological competition. In that sense, resistance itself is not new. What is new—and arguably more concerning—is the scale and intensity at which resistance genes are now being introduced and mobilized through human activity. The addition of clinically relevant antibiotics and antibiotic-resistant bacteria (ARB) into soil environments effectively transforms these ecosystems into what some researchers have described as “genetic reactors,” where selective pressures favor the proliferation and exchange of resistance traits (Ahmad et al., 2026; Wu et al., 2025).
Under such conditions, horizontal gene transfer (HGT) becomes a central mechanism. Mobile genetic elements facilitate the movement of resistance genes across taxonomic boundaries, potentially accelerating the spread of antimicrobial resistance within soil microbiomes and beyond (Jadeja & Worrich, 2022). The consequences are not confined to microbial ecology alone; they extend to ecosystem functionality. For instance, elevated antibiotic concentrations have been associated with disruptions in nitrogen cycling, particularly through the suppression of nitrifying microorganisms, which may, in turn, reduce soil nitrogen-processing efficiency (Yan et al., 2026).
3.1.3 Soil Fauna as Mediators of Resistance Dynamics
An additional layer of complexity emerges when considering the role of soil fauna. Organisms such as earthworms and collembolans, often overlooked in discussions of AMR, harbor their own microbiomes and interact intimately with surrounding microbial communities. Interestingly, their role appears to be somewhat dualistic. On one hand, these organisms may act as vectors, facilitating the movement of resistance genes through soil layers and food webs. On the other, there is growing evidence that certain fauna—particularly earthworms—may exert a mitigating effect. Their gut environments can selectively suppress or degrade antibiotic-resistant bacteria, suggesting a form of biological filtration that may reduce the overall burden of resistance determinants in soil systems (Jadeja & Worrich, 2022; Phan et al., 2024).
3.1.4 Implications for One Health and Sustainable Soil Management
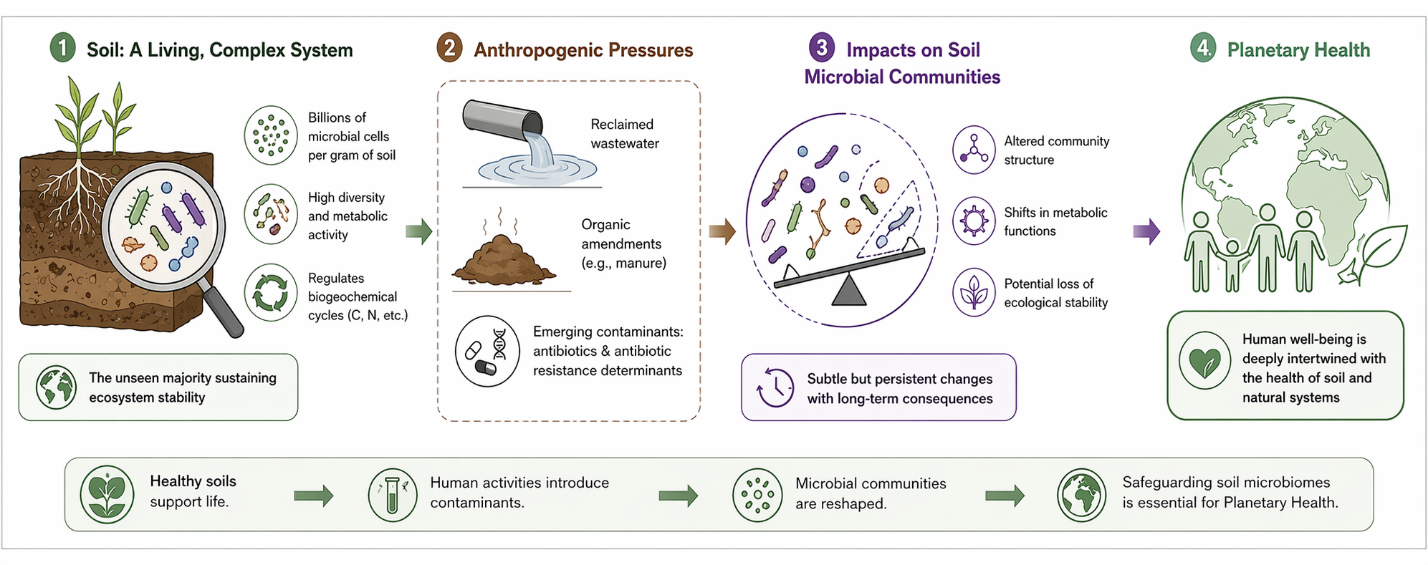
Figure 1: Hidden Resistome in Rice Paddies: How Reclaimed Water Irrigation Reshapes Soil Microbiomes and Amplifies AMR Risks. Soil is a dynamic, microbially rich ecosystem where reclaimed wastewater and agricultural inputs introduce antibiotics and resistance genes, subtly reshaping microbial structure and function. These interactions across the soil–plant–water continuum position rice paddies as critical nodes in the environmental dissemination of antimicrobial resistance, linking ecosystem integrity to human health.
Table 1. Effects of Irrigation Water Quality and Fertilization Regimes on Soil Resistome Dynamics in Agroecosystems. This table summarizes how reclaimed wastewater irrigation and different fertilization strategies influence ARG abundance and mobile genetic elements (MGEs) in soil systems. It highlights the dual role of nutrient inputs and irrigation sources in either suppressing or amplifying antimicrobial resistance within rhizosphere and bulk soils.
|
Reference
|
System/Matrix
|
Water Source
|
Fertilizer
|
Main ARG Class
|
Abundance Shift
|
MGE Status
|
Key Ecological Insight
|
|
Cui et al. (2022)
|
Tomato rhizosphere
|
Reclaimed wastewater (RWW)
|
N (NaNO₃)
|
Multidrug
|
24.06–73.09% decrease
|
Increased vs control
|
Nitrogen regulation reduces ARG burden in soil
|
|
Cui et al. (2022)
|
Tomato rhizosphere
|
RWW
|
P (CaMgO₄P⁺)
|
Aminoglycoside
|
50.13–58.80% decrease
|
Variable
|
Soil pH and total N drive ARG variation
|
|
Cui et al. (2022)
|
Tomato bulk soil
|
RWW
|
P fertilizer
|
Sulfonamide
|
Up to 35.84% increase
|
Positive correlation
|
P fertilization may enrich ARGs in bulk soil
|
|
Cui et al. (2022)
|
Tomato rhizosphere
|
RWW
|
K (K₂SO₄)
|
Tetracycline
|
46.52–53.64% decrease
|
Moderate increase
|
K₂SO₄ more effective than KCl
|
|
Xu et al. (2024)
|
Paddy soil
|
Paddy water
|
Swine compost
|
MLSB / Multidrug
|
Significant enrichment
|
Marker genes detected
|
Compost increases resistome load
|
|
Xu et al. (2024)
|
Rice roots
|
Paddy water
|
Swine compost
|
Beta-lactam
|
Substantial presence
|
intI1 / tnpA detected
|
Elevated HGT from soil to plant
|
|
Phan et al. (2024)
|
Spinach soil
|
Treated wastewater (TMW)
|
Antibiotic mixture
|
Multidrug
|
Highly diverse and elevated
|
9 MGEs detected
|
Residual antibiotics increase ARG diversity
|
|
Phan et al. (2024)
|
Radish rhizosphere
|
TMW
|
Antibiotic mixture
|
FCA resistance
|
Lower than bulk soil
|
Reduced HGT
|
Rhizosphere resilience moderates ARG spread
|
|
Della-Negra et al. (2026)
|
Loamy clay soil
|
Treated wastewater (TWW)
|
None
|
sul1
|
Significant increase
|
intI1 enrichment
|
Long-term TWW promotes ARG accumulation
|
|
Della-Negra et al. (2026)
|
Lettuce roots
|
Raw wastewater
|
None
|
ermB
|
Selective modulation
|
Barrier effect
|
Root microbiome limits AMR transfer
|
Taken together, these dynamics underscore the necessity of adopting a One Health perspective when addressing antimicrobial resistance in soil ecosystems. The soil–plant–water continuum is not an isolated system; rather, it functions as a conduit through which resistance can move between environmental reservoirs and human populations. Managing soil health, therefore, extends beyond agronomic productivity—it becomes a matter of public health relevance. Strategies such as precision fertilization, improved wastewater treatment technologies, and controlled reuse practices may offer pathways to mitigate these risks. Still, the challenge lies in balancing agricultural sustainability with the preservation of microbial integrity, ensuring that soil remains a resilient and functional component of the Earth system rather than a persistent source of resistance dissemination (EFSA Panel on Biological Hazards, 2021; Ahmad et al., 2026).
3.2 Soil as a Source of Bioactive Compounds in the Context of Soil Biota, AMR, and Planetary Health
3.2.1 Soil Biodiversity as a Reservoir of Bioactive Potential
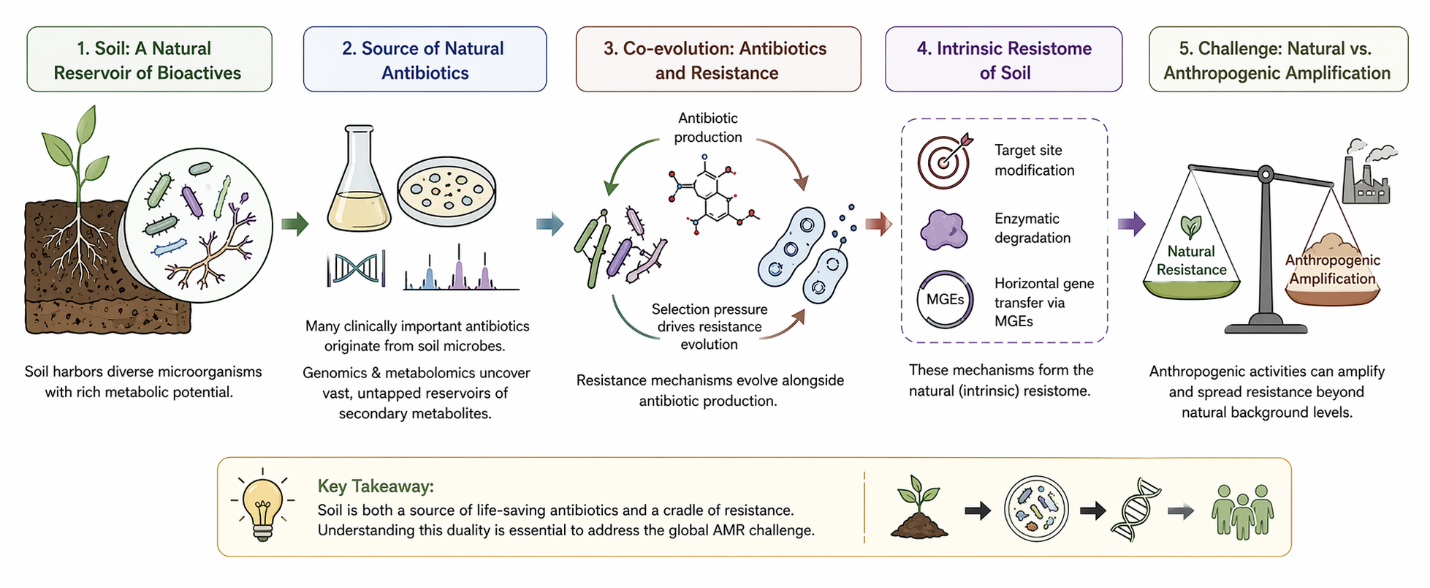
It is tempting—perhaps too tempting—to think of soil merely as a substrate for plant growth. Yet, when examined more closely, it becomes evident that soil functions as a remarkably dynamic biochemical reservoir. Within its microscale heterogeneity exists a dense and metabolically versatile microbial community capable of producing an extraordinary diversity of bioactive compounds, many of which have historically underpinned antibiotic discovery. This immense biochemical potential, however, is not isolated from broader ecological processes. Rather, it is embedded within a system that regulates nutrient cycling, supports plant productivity, and ultimately sustains planetary health (Zhu et al., 2019; Ondon et al., 2021).
At the same time, this reservoir is increasingly influenced by anthropogenic pressures. The introduction of reclaimed wastewater, agricultural runoff, and manure amendments alters not only nutrient availability but also the chemical landscape of soil environments. These inputs may unintentionally reshape microbial metabolic pathways, influencing both the production and degradation of bioactive compounds. In this sense, soil is no longer just a source of beneficial metabolites—it is becoming a contested space where ecological function and anthropogenic disturbance intersect (Phan et al., 2024; Wu et al., 2025).
3.2.2 Bioactive Compounds, Resistome Evolution, and Selective Pressures
There is, however, an inherent paradox in the role of soil-derived bioactive compounds. Many antibiotics originate from soil microorganisms, functioning naturally as signaling molecules or competitive tools within microbial communities. This long evolutionary history has given rise to what is often described as the intrinsic resistome—a baseline level of resistance shaped by ecological interactions over geological timescales. Yet, the rapid influx of synthetic and clinical antibiotics into soil systems has disrupted this balance, amplifying resistance far beyond its natural context (Ahmad et al., 2026; Selvarajan et al., 2023).
Under these altered conditions, selective pressures intensify. Antibiotics, heavy metals, and other co-contaminants collectively drive the proliferation of resistance genes, often co-located on mobile genetic elements. Horizontal gene transfer (HGT) then facilitates their spread across diverse microbial taxa, effectively accelerating the evolution of the resistome. This process, while mechanistically fascinating, raises concerns about the long-term sustainability of soil ecosystems and their capacity to maintain essential functions (Jadeja & Worrich, 2022; EFSA Panel on Biological Hazards, 2021).
3.2.3 Agricultural Practices and the Modulation of Soil Bioactivity
Agricultural management practices further complicate this picture. Fertilization strategies, particularly the application of nitrogen, phosphorus, and potassium, do not merely influence crop yield—they also modulate microbial community structure and activity. Some evidence suggests that targeted fertilization approaches may mitigate the accumulation of antibiotic resistance genes in reclaimed water–irrigated soils, although the mechanisms remain incompletely understood (Cui et al., 2022). This introduces an important, if somewhat underexplored, possibility: that soil management practices could be strategically designed to preserve beneficial bioactivity while limiting the spread of resistance.
3.2.4 Toward Integrated Soil Stewardship
Ultimately, the role of soil as a source of bioactive compounds cannot be disentangled from its role in antimicrobial resistance dynamics. The same microbial diversity that offers solutions—new antibiotics, novel metabolites—also harbors the capacity for resistance proliferation under selective pressure. This duality reinforces the need for a One Health perspective, where soil stewardship is recognized as integral to both environmental sustainability and global health. Managing this balance, admittedly, is not straightforward. It requires coordinated efforts across disciplines to ensure that soil remains a source of innovation rather than a reservoir of risk (EFSA Panel on Biological Hazards, 2021; Ahmad et al., 2026).
3.3 Antimicrobial Resistance (AMR) Reservoirs in the Context of Soil Biota and Planetary Health
3.3.1 Soil as a Dynamic Reservoir of Resistance
Soil, when viewed beyond its agronomic function, reveals itself as a remarkably dense and dynamic microbial habitat—arguably the most complex on Earth. Within this microcosm, vast and diverse microbial communities regulate essential biogeochemical cycles, quietly sustaining planetary habitability. It is precisely this richness, however, that makes soil both resilient and vulnerable. As agricultural systems increasingly rely on reclaimed wastewater and organic amendments, a subtle yet consequential shift is occurring: soil is becoming an active reservoir for antimicrobial resistance (AMR), rather than merely a passive recipient (Zhu et al., 2019; Ondon et al., 2021).
3.3.2 From Natural Resistome to Anthropogenic Amplification
There is, of course, a long-standing natural resistome within soil ecosystems—an evolutionary legacy shaped by microbial competition over millions of years. Yet, the introduction of anthropogenic pressures, particularly antibiotics and antibiotic-resistant bacteria (ARB), appears to have intensified this baseline in ways that are not entirely predictable. Under such conditions, soil environments may function as “genetic reactors,” where selective pressures and dense microbial interactions promote horizontal gene transfer (HGT) via mobile genetic elements such as plasmids and integrons (Ahmad et al., 2026; Phan et al., 2024). The extent of this transformation remains somewhat contested. While some studies report minimal disruption due to competitive microbial dynamics, others suggest that prolonged reclaimed wastewater irrigation can substantially enrich the soil resistome, in some cases by several orders of magnitude (Della-Negra et al., 2026).
3.3.3 Biological Mediators and Ecological Consequences
Adding further complexity, soil fauna contribute in ways that are not entirely linear. Organisms such as collembolans may facilitate the movement of resistance genes across trophic levels, effectively acting as vectors. Conversely, earthworms—perhaps unexpectedly—may exert a mitigating influence. Their gut-associated microbiomes have been shown to suppress certain antibiotic-resistant bacteria, suggesting a form of biological filtration that could reduce resistance burdens under specific conditions (Ondon et al., 2021; Phan et al., 2024).
3.3.4 Implications for One Health and Ecosystem Function
These intertwined processes ultimately reinforce the need for a One Health perspective. The soil–plant–water continuum represents not just an environmental interface but a pathway through which resistance can circulate across ecological and human systems. Moreover, antibiotic residues may disrupt key ecosystem functions, including nitrogen cycling, potentially diminishing soil nutrient efficiency (Yan et al., 2026). Addressing these challenges will require carefully balanced strategies—improving wastewater treatment, refining fertilization practices, and preserving microbial integrity—to ensure that soil remains a stabilizing force rather than a persistent reservoir of resistance (Ahmad et al., 2026; Zhu et al., 2019).
3.4 Planetary Health Implications in the Context of Soil Biota and Antimicrobial Resistance
3.4.1 Soil Integrity as a Pillar of Planetary Health
It is easy—perhaps misleadingly so—to reduce soil to its agricultural function, to think of it primarily as a medium that supports plant growth. Yet soil is better understood as a living, breathing interface between biological, chemical, and physical systems. Within a single gram exists an immense diversity of microorganisms whose collective activity regulates fundamental planetary processes, including carbon sequestration and nitrogen transformation. In this sense, soil does not merely support life—it helps sustain the conditions that make life possible. This perspective aligns closely with the concept of Planetary Health, where human well-being is inseparable from the stability of natural ecosystems (Zhu et al., 2019; EFSA Panel on Biological Hazards, 2021).
However, this equilibrium appears increasingly fragile. The widespread use of reclaimed wastewater (RWW) and organic waste in agriculture introduces a complex mixture of antibiotics, resistant microorganisms, and chemical stressors into soil systems. While these practices are often promoted as sustainable solutions to water scarcity, they may simultaneously impose selective pressures that reshape microbial communities in ways that are not fully predictable (Phan et al., 2024; Wu et al., 2025).
3.4.2 Anthropogenic Pressures and Resistome Expansion
There is, undeniably, a natural baseline of resistance within soil ecosystems—the intrinsic resistome—formed through long evolutionary processes. Yet, the rapid influx of anthropogenic contaminants has shifted this balance. Soil environments, under these conditions, begin to resemble “genetic reactors,” where high microbial density and persistent selective pressures facilitate horizontal gene transfer (HGT) via mobile genetic elements such as plasmids and integrons (Ahmad et al., 2026; Ondon et al., 2021). This amplification of resistance is not merely a microbiological concern; it has broader ecological consequences.
For instance, antibiotic residues can interfere with microbial-mediated nutrient cycling, particularly nitrogen transformations. Disruptions to nitrification and denitrification processes may reduce soil nutrient efficiency, with cascading effects on plant productivity and ecosystem resilience (Yan et al., 2026). At a planetary scale, even modest perturbations in these cycles could accumulate, influencing broader biogeochemical stability.
3.4.3 Biological Intermediaries and Ecological Feedbacks
The role of soil fauna introduces an additional layer of complexity that is, perhaps, still underappreciated. Organisms such as collembolans and earthworms do not simply inhabit soil—they actively reshape it, both physically and biologically. Some fauna may inadvertently facilitate the spread of antimicrobial resistance by acting as vectors, transferring resistance genes through trophic interactions. Others, however, appear to contribute to mitigation. Earthworms, for example, have been observed to reduce the abundance of antibiotic-resistant bacteria through gut-associated microbial processes, suggesting a form of natural attenuation that may partially counterbalance anthropogenic inputs (Jadeja & Worrich, 2022; Della-Negra et al., 2026).
At the same time, abiotic stressors—such as heavy metals and residual agrochemicals—interact with biological processes, reinforcing selective pressures that sustain resistance traits. These co-selection mechanisms further complicate efforts to disentangle cause and effect within soil systems (Rad et al., 2022).
3.4.4 Toward a One Health Framework for Soil Stewardship
Taken together, these dynamics point toward a broader realization: soil is not an isolated component of the environment but a central node within the soil–plant–water–human continuum. The dissemination of antimicrobial resistance through this system underscores the urgency of adopting a One Health framework, where environmental management is directly linked to public health outcomes. Managing soil health, therefore, becomes more than an agronomic priority—it is a global imperative.
Moving forward, the challenge lies in balancing sustainability with precaution. Strategies such as advanced wastewater treatment, precision fertilization, and improved monitoring of resistance determinants may offer viable pathways. Yet, these approaches must be implemented with an awareness of ecological complexity, ensuring that interventions do not inadvertently exacerbate the very problems they aim to solve. In this context, preserving soil integrity is not simply about maintaining productivity—it is about safeguarding the ecological foundations upon which planetary health ultimately depends (Ahmad et al., 2026; Zhu et al., 2019).
3.5 Management Strategies and Future Research in the Context of Soil Biota, AMR, and Planetary Health
3.5.1 Reframing Soil Management Beyond Productivity
Soil, despite its frequent treatment as a static agricultural input, is perhaps better understood as a dynamic, living interface—one that mediates interactions between biological communities, chemical processes, and human interventions. Within this system, microbial diversity underpins essential functions such as nutrient cycling and organic matter turnover, quietly sustaining agricultural productivity and broader ecosystem stability. Yet, as reclaimed wastewater (RWW) and organic amendments become more widely integrated into farming systems, particularly in water-intensive landscapes, this balance appears increasingly fragile. The introduction of antibiotics and antibiotic-resistant bacteria (ARB) into soil systems is not merely an environmental side effect; it is, increasingly, a defining challenge for sustainable land management (Zhu et al., 2019; Phan et al., 2024).
3.5.2 Mitigating Resistome Expansion Through Targeted Interventions
At the heart of this challenge lies the transition from the intrinsic resistome—long embedded within soil microbial ecology—to an anthropogenically amplified resistome shaped by continuous external inputs. Under these conditions, soil environments may function as “genetic reactors,” where selective pressures promote horizontal gene transfer (HGT) via mobile genetic elements such as plasmids and integrons (Ahmad et al., 2026; Ondon et al., 2021). Addressing this requires interventions that are, admittedly, both technically robust and ecologically sensitive.
Improving wastewater treatment technologies is often presented as a primary solution. Advanced treatment processes, including membrane filtration and advanced oxidation, have shown promise in reducing antibiotic residues and resistance determinants before irrigation use. However, their implementation remains uneven, particularly in regions where water reuse is most critical (Duarte et al., 2022; Wu et al., 2025). Complementary to this, precision fertilization strategies—especially those optimizing nitrogen inputs—may help regulate microbial activity and limit the proliferation of antibiotic resistance genes (ARGs), although the mechanisms are not yet fully resolved (Rad et al., 2022; Yan et al., 2026).
3.5.3 Biological Mediation and Nature-Based Solutions
There is also growing interest in biological approaches to mitigation, though these remain, in some respects, exploratory. Soil fauna, particularly earthworms, appear to play a role that is more complex than previously assumed. While certain organisms may facilitate the spread of resistance through trophic interactions, others may contribute to attenuation. Earthworms, for example, have been observed to reduce the abundance of ARB through gut-associated microbial processes, suggesting a potential avenue for natural bioremediation (Jadeja & Worrich, 2022; Della-Negra et al., 2026). Whether such processes can be reliably harnessed at scale remains an open question.
3.5.4 Future Research Directions and One Health Integration
Looking forward, it becomes evident that addressing AMR in soil systems requires more than isolated technical fixes. There is a need for integrated research frameworks that link soil microbiology, agricultural practices, and public health outcomes. Long-term field studies, in particular, are essential to capture the cumulative effects of reclaimed water irrigation and management strategies on soil resistomes (Selvarajan et al., 2023; Phan et al., 2024). Additionally, improved monitoring tools—capable of tracking ARG dynamics across the soil–plant–water continuum—will be critical for risk assessment and policy development.
Ultimately, the management of soil in the age of antimicrobial resistance demands a One Health perspective. Soil health is no longer solely an agronomic concern; it is a shared environmental and public health responsibility. Balancing productivity with ecological integrity will require coordinated efforts, informed by both scientific insight and practical feasibility, to ensure that soil remains a resilient system rather than a persistent reservoir of resistance (EFSA Panel on Biological Hazards, 2021; Ahmad et al., 2026).