1. Introduction
Antimicrobial resistance (AMR) has gradually evolved from a largely clinical concern into a broader global health emergency that now intersects with microbiology, public health, biotechnology, and even socioeconomic stability. What once appeared manageable through successive generations of antibiotics is increasingly becoming unpredictable, particularly as resistant pathogens continue to emerge faster than many conventional therapeutics can be developed. The problem is no longer confined to isolated hospital-acquired infections; rather, resistant microorganisms are now deeply embedded within community settings, environmental reservoirs, and complex host-associated microbial ecosystems (Mantravadi et al., 2019). Although antibiotics transformed medicine throughout the twentieth century, their prolonged and frequently indiscriminate use has accelerated adaptive microbial responses, ultimately reduced therapeutic effectiveness and narrowed available treatment options (Coates et al., 2002; Jubeh et al., 2020).
For decades, antimicrobial discovery relied heavily on targeting a relatively small number of bacterial processes, particularly DNA replication, protein synthesis, and cell wall biosynthesis. Initially, these approaches were remarkably successful. Yet bacteria, perhaps unsurprisingly, have demonstrated extraordinary evolutionary flexibility. Resistance mechanisms such as efflux pumps, enzymatic degradation, target modification, and horizontal gene transfer have become increasingly widespread, complicating infection management in both clinical and environmental contexts (Mantravadi et al., 2019). Consequently, antimicrobial research has shifted toward identifying more selective and less resistance-prone molecular targets. Particular attention has been directed toward bacterial metabolic pathways absent in mammalian hosts, as these systems may provide opportunities for narrow-spectrum therapies with reduced off-target toxicity (Hudson et al., 2008).
Among the most promising targets are enzymes involved in the shikimate pathway and the L,L-diaminopimelate biosynthesis pathway. These metabolic systems are essential for bacterial survival yet absent in humans, making them attractive candidates for antimicrobial intervention (Gonzalez-Bello, 2016; Hudson et al., 2008). Similarly, structurally conserved cell wall intermediates such as Lipid II have gained considerable attention because their functional conservation reduces the likelihood of rapid mutational escape. The discovery of teixobactin represented an especially important milestone in this regard, not necessarily because it completely solved resistance-associated challenges, but because it demonstrated that novel antimicrobial scaffolds can still emerge from previously inaccessible microbial populations (Ling et al., 2015). Compounds such as lassomycin further reinforce the growing interest in targeting bacterial proteostasis systems, particularly ATP-dependent proteases associated with persistence and survival under stress conditions (Gavrish et al., 2014).
At the same time, it has become increasingly evident that antimicrobial resistance cannot be understood solely through the lens of planktonic bacterial growth. Biofilms, which are structured microbial communities embedded within extracellular polymeric matrices, fundamentally alter microbial physiology and antimicrobial susceptibility. These highly organized systems create protective microenvironments that shield bacteria from antibiotics, immune responses, and environmental stressors (Brackman & Coenye, 2015). In many chronic infections, including oral, pulmonary, and device-associated infections, biofilm formation appears to contribute substantially to treatment failure and recurrence. Interestingly, the resilience of biofilms is not simply structural; it is also regulatory. Quorum sensing (QS), the bacterial communication process responsible for coordinating collective behavior, plays a central role in biofilm maturation, virulence expression, and microbial adaptation (Gutierrez et al., 2009; Han & Lu, 2009).
Because of this, quorum sensing inhibition has emerged as a particularly intriguing “anti-virulence” strategy. Rather than directly killing bacterial cells, quorum sensing inhibitors (QSIs) attempt to disarm pathogenic behaviors by disrupting microbial communication networks (Brackman & Coenye, 2015). This distinction may prove important, as therapies that reduce virulence without exerting strong bactericidal pressure could theoretically slow the emergence of resistance. Several compounds targeting LuxS, MTAN, and related signaling systems have demonstrated encouraging activity against biofilm-associated pathogens (Gutierrez et al., 2009; Lee et al., 2005). Likewise, azithromycin and other signaling-modulating agents have shown the capacity to interfere with quorum sensing-regulated alginate production in Pseudomonas aeruginosa, thereby weakening biofilm integrity and persistence (Hoffmann et al., 2007). Natural compounds capable of reducing intracellular cyclic-di-GMP levels, such as raffinose-derived molecules, have also demonstrated anti-biofilm potential, suggesting that microbial signaling pathways may offer multiple intervention points beyond traditional antibiotic targets (Kim et al., 2016).
Another challenge complicating antimicrobial therapy is the existence of persister cells. These dormant or metabolically inactive bacterial populations are not necessarily genetically resistant, yet they tolerate antibiotic exposure and frequently contribute to recurrent infections. Persisters are particularly problematic in chronic diseases where prolonged treatment fails to fully eradicate microbial reservoirs. Recent work involving Clp proteases suggests that manipulating bacterial proteolytic systems may provide a viable strategy for eliminating these difficult-to-target populations (Conlon et al., 2013; Gavrish et al., 2014). Although still emerging, this field reflects a broader shift in antimicrobial research away from simple growth inhibition toward a deeper understanding of microbial physiology, stress adaptation, and persistence mechanisms.
Parallel to these molecular developments, advances in genomics and computational biology have profoundly reshaped antimicrobial discovery pipelines. Traditional “culture-and-screen” approaches, while historically productive, overlooked the overwhelming majority of microbial diversity because many microorganisms remain difficult or impossible to cultivate under conventional laboratory conditions. Genome mining platforms such as antiSMASH now allow researchers to identify cryptic biosynthetic gene clusters that may encode previously unknown antimicrobial metabolites (Blin et al., 2017). Meanwhile, culturomics and metagenomics have expanded access to microbial communities that were once considered inaccessible, revealing an extraordinary reservoir of metabolic potential within environmental and host-associated microbiota (Lagier et al., 2018).
These technological advances are especially relevant in microbiome research, where the distinction between pathogenicity and ecological imbalance is increasingly blurred. Dysbiosis, rather than single-pathogen infection alone, is now recognized as an important contributor to oral, gastrointestinal, vaginal, and systemic diseases (Giordano-Kelhoffer et al., 2022). Oral microbial imbalance, for example, has been associated not only with periodontal disease but also with broader inflammatory and neurodegenerative conditions, including Alzheimer’s disease (Borsa et al., 2021). This evolving perspective has encouraged the development of microbiome-centered therapeutics aimed not merely at eradicating pathogens but at restoring ecological equilibrium.
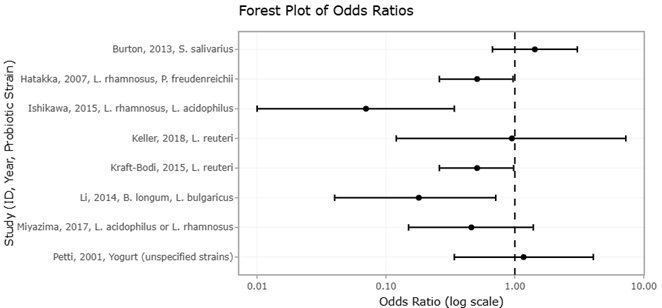
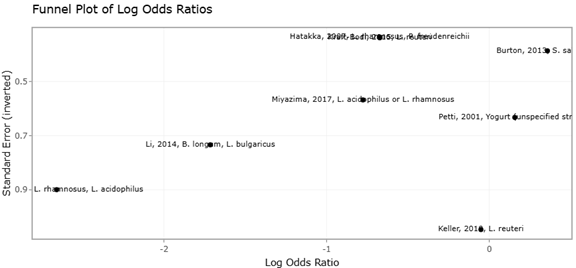
In this context, probiotics and next-generation probiotics (NGPs) have emerged as particularly promising therapeutic tools. Unlike traditional antimicrobials, probiotics may exert beneficial effects through competitive exclusion, immunomodulation, metabolite production, and stabilization of microbial communities (Abouelela & Helmy, 2024). Clinical investigations involving Lactobacillus, Bifidobacterium, and Streptococcus salivarius strains have demonstrated reductions in oral Candida colonization and improvements in microbial balance across several patient populations (Burton et al., 2013; Hatakka et al., 2007; Ishikawa et al., 2015). Additional studies involving recurrent candidiasis and bacterial vaginosis have suggested that probiotic formulations may also influence vaginal microbial ecosystems and inflammatory responses (Bisanz et al., 2014; Bradshaw et al., 2012; Mastromarino et al., 2009; Oerlemans et al., 2020).
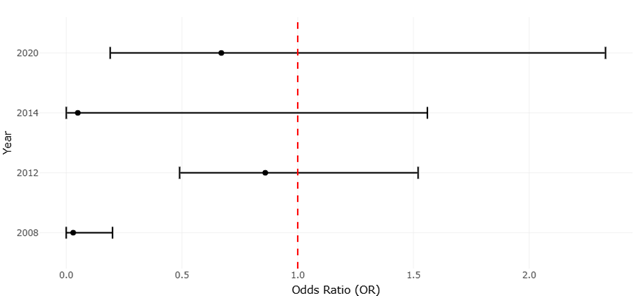
Still, probiotic efficacy is not always uniform. Outcomes often vary according to strain composition, dosing regimen, microbial baseline composition, and host physiology. Multi-strain formulations frequently demonstrate stronger or more consistent effects than single-strain preparations, possibly because of synergistic microbial interactions and broader ecological coverage (Kraft-Bodi et al., 2015; Li et al., 2014; Miyazima et al., 2017). Likewise, probiotic-enriched dietary interventions, including fermented dairy products, have shown measurable influence on oral microbial composition and pathogen suppression (Keller & Kragelund, 2018; Petti et al., 2001). These findings suggest that microbiome modulation may become an increasingly important complement to conventional antimicrobial therapy, particularly in conditions associated with chronic dysbiosis and biofilm persistence.
Natural products also continue to play an indispensable role in antimicrobial discovery. Essential oils and phytochemical-derived compounds possess diverse antibacterial, antifungal, and immunomodulatory activities, often acting through multiple mechanisms simultaneously (Freires et al., 2015). This mechanistic diversity may reduce the likelihood of rapid resistance development compared with highly specific single-target drugs. Importantly, the integration of natural compounds, biofilm-disrupting agents, probiotics, and precision molecular inhibitors reflects a broader conceptual transition within antimicrobial science. The field is gradually moving away from a singular “kill-the-pathogen” philosophy toward more ecologically informed and systems-oriented therapeutic strategies.
Taken together, contemporary antimicrobial research appears to be entering a far more integrative phase—one that combines microbiome science, synthetic biology, genomics, bioinformatics, and molecular pharmacology to address the growing complexity of infectious disease management. Although no single intervention is likely to resolve the AMR crisis independently, the convergence of targeted molecular therapies, biofilm disruption strategies, microbiome restoration, and natural product discovery offers a more adaptive and potentially sustainable framework for future antimicrobial development.