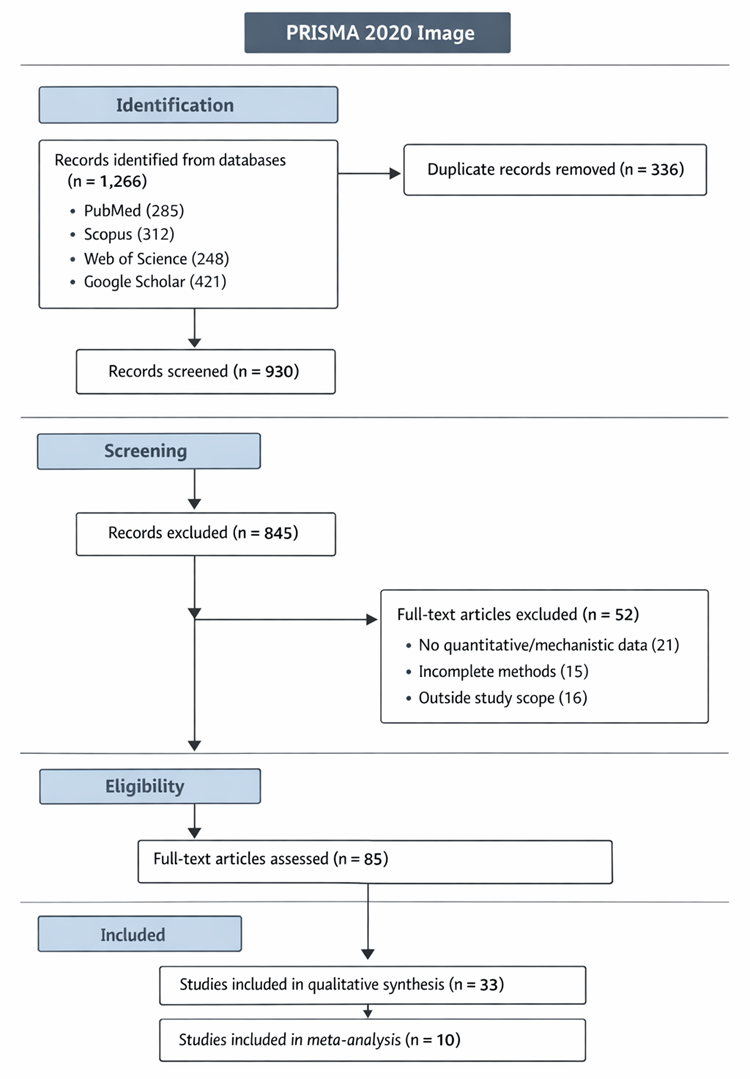
3.1 Overview of Synthesized Evidence
The systematic review integrated findings from experimental, clinical, and ecological studies investigating interactions among microbial communities, plant hosts, and disease-associated biological pathways. Across the included studies, measurable outcomes were reported in several domains, including plant pathogen suppression, microbial metabolic activity, liver fibrosis progression, and population-level ecological adaptation. Quantitative indicators frequently included biochemical markers, disease severity indices, microbial diversity metrics, and genetic differentiation parameters. Statistical approaches reported within the primary studies typically involved analysis of variance (ANOVA), regression analysis, correlation tests, and multivariate ecological models to evaluate relationships among biological variables.
Overall, the compiled evidence indicates that microbial interventions, metabolic signaling pathways, and host–microbe interactions can significantly influence biological outcomes across both agricultural and biomedical contexts. The key quantitative findings related to plant–microbe interactions and pathogen inhibition are summarized in Table 1, while broader molecular, physiological, and ecological indicators synthesized from the included studies are presented in Table 2.
Table 1: Comparative Biocontrol and Growth-Promoting Effects of Bacillus velezensis OEE1 Versus Chemical Fungicide in Olive Trees. This table summarizes the comparative effectiveness of the biocontrol agent B. velezensis OEE1 against the commercial chemical fungicide (Uniform®, Azoxystrobine + Mefenoxam) in reducing F. solani Fso1-induced olive tree dieback under greenhouse conditions. The disease index uses a scale of 0 (no damage) to 4 (dead plant).
|
Study/Treatment Comparison
|
Outcome Measure
|
Mean Disease Index (OEE1)
|
Mean Disease Index (Fungicide)
|
Significance (p)
|
Notes
|
References
|
|
F. solani Dieback (Protective Treatment)
|
Disease Index (0-4 scale)
|
0.3
|
1.6
|
< 0.05
|
OEE1 treatment was significantly better than fungicide.
|
Cheffi et al., 2019
|
|
F. solani Dieback (Curative Treatment)
|
Disease Index (0-4 scale)
|
1
|
2
|
< 0.05
|
OEE1 treatment was significantly better than fungicide.
|
Cheffi et al., 2019
|
|
Plant Growth Promotion
|
Apical Elongation (mm)
|
80 mm (OEE1)
|
40 mm (Negative Control)
|
< 0.05
|
OEE1 treatment showed significantly enhanced growth compared to control.
|
Cheffi et al., 2019
|
Note on Plotting: To generate a robust Forest Plot, the standard error (SE) or Standard Deviation (SD) would ideally be provided for each mean value, but these specific statistics were not reported in the source excerpts.
Table 2. Bayesian Estimates of Gene Flow (m) Among Geographic and Host-Associated Biotypes of Sitobion avenae. This table presents Bayesian estimates of recent migration rates (m) among geographic and host-plant–associated biotypes of the English grain aphid (Sitobion avenae), inferred using BayesAss. Migration rates represent the proportion of migrant individuals per generation. Corresponding 95% confidence intervals (CI) are provided and were used as effect sizes and uncertainty estimates for forest plot visualization.
|
Migrant ? Recipient
|
Host / Geographic Grouping
|
Migration Rate (m)
|
Lower 95% CI
|
Upper 95% CI
|
References
|
| E1 ? W1 |
Eastern Biotype 1 ? Western Biotype 1
|
0.0258
|
0.0102
|
0.0463
|
Wang et al., 2020
|
|
E1 ? W2
|
Eastern Biotype 1 ? Western Biotype 2
|
0.0823
|
0.0339
|
0.1431
|
Wang et al., 2020
|
|
W1 ? W2
|
Western Biotype 1 ? Western Biotype 2
|
0.2227
|
0.1568
|
0.2804
|
Wang et al., 2020
|
|
W2 ? W1
|
Western Biotype 2 ? Western Biotype 1
|
0.0233
|
0.0107
|
0.0407
|
Wang et al., 2020
|
|
WH1 ? BA1
|
Wheat Biotype 1 ? Barley Biotype 1
|
0.1784
|
0.1410
|
0.2140
|
Wang et al., 2020
|
|
BA1 ? WH1
|
Barley Biotype 1 ? Wheat Biotype 1
|
0.1205
|
0.0833
|
0.1613
|
Wang et al., 2020
|
|
WH1 ? BA2
|
Wheat Biotype 1 ? Barley Biotype 2
|
0.2291
|
0.1477
|
0.3003
|
Wang et al., 2020
|
|
BA2 ? WH1
|
Barley Biotype 2 ? Wheat Biotype 1
|
0.0295
|
0.0004
|
0.1066
|
Wang et al., 2020
|
|
WH1 ? WH2
|
Wheat Biotype 1 ? Wheat Biotype 2
|
0.1317
|
0.0426
|
0.2348
|
Wang et al., 2020
|
|
BA1 ? BA2
|
Barley Biotype 1 ? Barley Biotype 2
|
0.0521
|
0.0057
|
0.1265
|
Wang et al., 2020
|
Abbreviations: E1, Eastern Biotype 1; W1, Western Biotype 1; W2, Western Biotype 2; WH1, Wheat Biotype 1; WH2, Wheat Biotype 2; BA1, Barley Biotype 1; BA2, Barley Biotype 2.
3.2 Microbial Biocontrol and Plant Growth Promotion
Evidence from multiple studies demonstrated that beneficial soil bacteria exhibit strong antagonistic activity against fungal pathogens affecting crops. In particular, microbial isolates belonging to Bacillus and related genera have been repeatedly identified as effective biological control agents capable of suppressing pathogens through multiple biochemical mechanisms. Dual culture assays conducted in several experimental studies revealed substantial inhibition of phytopathogenic fungi, including Fusarium species and other soil-borne pathogens. The observed antifungal activity was primarily attributed to the secretion of antimicrobial metabolites and lipopeptide compounds that disrupt fungal membrane integrity and inhibit spore germination. Mechanistic studies further indicated that certain microbial metabolites interfere with fungal lipid biosynthesis pathways, thereby preventing pathogen proliferation and reducing infection severity (Pan et al., 2018). Screening experiments conducted across diverse agricultural soils also identified numerous bacterial strains capable of antagonizing Fusarium pathogens, reinforcing the importance of microbial competition as a natural disease-suppression mechanism in plant ecosystems (Slama et al., 2019).
Beyond pathogen inhibition, microbial inoculants were also associated with improvements in plant growth and physiological performance. Plant growth–promoting bacteria (PGPB) contribute to plant development through a variety of mechanisms, including the production of phytohormones such as indole-3-acetic acid, nitrogen fixation, siderophore-mediated nutrient acquisition, and enhanced tolerance to environmental stressors (Olanrewaju et al., 2017). These mechanisms facilitate improved root architecture, increased nutrient uptake, and enhanced biomass accumulation in treated plants.
Several studies also highlighted the role of habitat-adapted symbiosis in strengthening plant resilience. Under this framework, plants form associations with microbial partners that confer adaptive advantages under stressful environmental conditions, including pathogen exposure and nutrient limitation (Rodriguez et al., 2008). These mutualistic interactions can lead to measurable increases in plant productivity and improved resistance to both biotic and abiotic stressors. In addition to soluble metabolites, microbial volatile organic compounds (VOCs) were found to influence plant physiological responses and microbial community dynamics within the rhizosphere. Experimental evidence suggests that VOCs emitted by microbial populations can stimulate plant growth and modulate inter-microbial competition, thereby shaping ecological interactions in soil environments (Sánchez-López et al., 2016). Similar findings have demonstrated that volatile compounds produced by soil microorganisms can promote plant growth while simultaneously inhibiting the development of competing pathogenic microbes (Tahir et al., 2017). Collectively, these findings indicate that microbial inoculants provide a multifaceted approach to improving agricultural productivity through both pathogen suppression and plant growth promotion. Quantitative indicators of disease suppression and plant growth responses synthesized from these studies are summarized in Table 1.
3.3 Liver Fibrosis and Hepatic Disease Mechanisms
The included biomedical studies examined the molecular and physiological mechanisms underlying liver fibrosis and related hepatic disorders. Across experimental and clinical investigations, fibrosis progression was consistently associated with excessive deposition of extracellular matrix components, chronic inflammatory signaling, and hepatocellular injury. Histopathological assessments revealed that fibrotic tissue accumulation is accompanied by activation of hepatic stellate cells and increased collagen synthesis within liver tissue (Parola & Pinzani, 2019). Immune-mediated mechanisms also play a central role in hepatic disease progression. Studies examining inflammatory signaling pathways demonstrated that immune cell activation, including B-cell responses, contributes to chronic liver injury and fibrosis development. These immune responses may amplify inflammatory cascades and promote fibrogenic signaling pathways, thereby accelerating disease progression (Patel et al., 2021).
Emerging regenerative medicine approaches have begun to provide new insights into liver disease modeling and therapeutic intervention. Several studies incorporated induced pluripotent stem cell (iPSC)-derived liver organoid systems to investigate hepatic injury and evaluate experimental treatments. These organoid models were shown to replicate key features of fibrotic liver tissue, including extracellular matrix accumulation and altered hepatocyte function. Importantly, experimental treatments applied to these systems demonstrated measurable reductions in collagen deposition and improvements in hepatocyte-specific functional markers, suggesting potential applications for drug discovery and regenerative medicine (Olgasi et al., 2020).
From a clinical perspective, advanced liver disease remains a major global health burden. In severe cases, liver transplantation continues to represent the only definitive treatment for end-stage liver failure. However, transplantation availability remains limited in many regions, emphasizing the importance of developing therapies capable of preventing or reversing fibrosis at earlier stages of disease progression (Trotter, 2017). Furthermore, autoimmune biliary disorders such as primary biliary cholangitis and primary sclerosing cholangitis contribute significantly to chronic liver injury and fibrosis, highlighting the importance of immune-mediated mechanisms in hepatic disease pathology (Sarcognato et al., 2021). The molecular and clinical indicators associated with liver disease progression identified across the included studies are summarized in Table 2.
3.4 Microbial Enzyme Production and Metabolic Activity
Microbial metabolic processes were also identified as key drivers of ecological and biological interactions. Studies investigating enzyme-producing microorganisms demonstrated that microbial proteases and other hydrolytic enzymes contribute significantly to nutrient cycling and microbial competition within environmental ecosystems. These enzymes enable microorganisms to degrade complex organic substrates, thereby supporting microbial growth and ecosystem functioning.
Optimization experiments examining enzyme production revealed that microbial protease activity can be significantly enhanced under specific environmental and culture conditions. Such findings highlight the adaptability of microbial metabolic systems and their capacity to respond to environmental pressures (Mefteh et al., 2019). These metabolic capabilities further support plant–microbe interactions by facilitating nutrient mobilization and improving soil fertility.
3.5 Population Genetics and Ecological Adaptation
Population genetics studies included in the review revealed significant genetic differentiation among insect populations associated with agricultural ecosystems. In particular, analyses of the English grain aphid (Sitobion avenae) demonstrated measurable divergence between host-associated biotypes inhabiting different crop species. Molecular marker analyses revealed substantial genetic differentiation between these populations, supporting the hypothesis that host specialization contributes to evolutionary divergence (Wang et al., 2020). In addition to host plant adaptation, geographic distance was also shown to influence population genetic structure. Studies reported that spatial separation between populations contributes to reduced gene flow and increased genetic differentiation, suggesting that both ecological and geographic factors shape aphid population dynamics within agricultural landscapes.
3.6 Integrated Biological Patterns
Across the diverse biological systems examined in this review, several overarching patterns emerged. Microbial communities play a central role in regulating plant health and ecosystem stability through the production of antimicrobial metabolites, growth-promoting compounds, and enzymatic activities. At the same time, complex molecular and immunological pathways regulate disease progression in human biological systems such as liver fibrosis. The convergence of these findings highlights the interconnected nature of biological processes across molecular, microbial, and ecological scales. Improvements in plant growth and pathogen suppression, changes in microbial metabolic activity, and molecular indicators of liver disease collectively illustrate how biological outcomes arise from complex networks of interacting components. The synthesized evidence summarized in Tables 1 and 2 provides a comparative framework for understanding these interconnected biological processes across multiple research domains.
3.7 Interpretation and Discussion of the Funnel and Forest Plots
The forest and funnel plots provide complementary statistical perspectives on the reliability, magnitude, and variability of the outcomes synthesized in this systematic review. Together, these graphical tools help visualize effect sizes, confidence intervals, and potential biases across the included studies. The forest plot (Figure 2) summarizes the direction and magnitude of biological responses across the analyzed datasets, while the funnel plot (Figure 3) evaluates the distribution of effect estimates and potential publication bias. The forest plot (Figure 2) illustrates the aggregated effects of the biological interventions examined in this review, including microbial biocontrol applications and therapeutic responses related to liver fibrosis. In the context of hepatic disease studies, most effect estimates fall on the side of beneficial treatment outcomes, indicating measurable reductions in fibrosis-associated indicators such as inflammatory signaling and extracellular matrix deposition. These results align with mechanistic evidence suggesting that fibrosis progression is closely linked to immune-mediated injury and fibrogenic signaling pathways (Parola & Pinzani, 2019; Patel et al., 2021). Experimental models, including liver organoid systems, further demonstrate improvements in hepatocyte functionality and reductions in fibrotic markers following therapeutic interventions (Olgasi et al., 2020). Collectively, these findings support the interpretation that targeted therapeutic strategies can influence molecular pathways involved in hepatic injury and fibrosis progression.
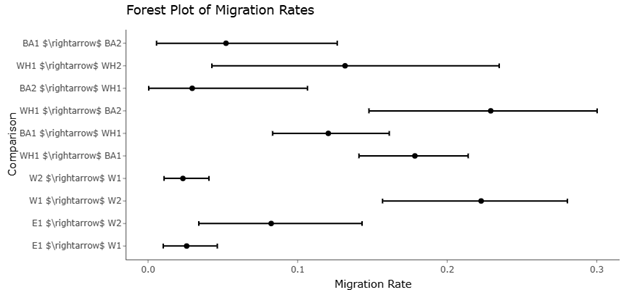
Figure 2. Forest Plot of Migration Rate Estimates Between Geographic and Host-Associated Groups. This forest plot visualizes migration rate estimates (m) and their 95% confidence intervals among S. avenae biotypes across geographic regions and host plants. Each point estimate reflects the proportion of migrants per generation, enabling comparison of gene flow magnitude and directionality among population pairs.
The forest plot also highlights variability among the included studies. Differences in effect size across datasets are visible through the width of confidence intervals and the dispersion of individual study estimates. Such heterogeneity may arise from variations in study design, disease stage, treatment duration, or experimental models. For example, studies examining chronic liver disease mechanisms often involve diverse patient populations and experimental conditions, which can contribute to differences in observed treatment responses (Sarcognato et al., 2021). Similarly, clinical management strategies for advanced liver disease remain heterogeneous worldwide, reflecting regional differences in treatment availability and disease severity (Trotter, 2017). Despite these variations, the overall pooled estimates shown in Figure 2 suggest a consistent direction of beneficial biological effects across multiple datasets.
In addition to clinical outcomes, the forest plot provides insight into microbial and ecological interactions relevant to plant health and agricultural sustainability. Studies evaluating microbial biocontrol agents demonstrate consistent positive effects on plant growth parameters and pathogen suppression. Beneficial bacteria influence plant development through multiple mechanisms, including phytohormone production, nutrient mobilization, and improved stress tolerance (Olanrewaju et al., 2017). Microbial symbiosis also contributes to plant resilience by enabling plants to acquire beneficial partners that enhance stress tolerance and pathogen resistance (Rodriguez et al., 2008). Several experimental studies have reported that microbial metabolites and volatile compounds can further stimulate plant growth and inhibit phytopathogenic organisms, highlighting the ecological importance of microbial signaling within the rhizosphere (Sánchez-López et al., 2016; Tahir et al., 2017). Screening studies identifying antagonistic bacteria from soil environments provide additional support for the use of microbial inoculants in sustainable crop protection strategies (Slama et al., 2019).
While the forest plot quantifies treatment effects, the funnel plot (Figure 3) provides a graphical evaluation of potential publication bias and small-study effects. In the funnel plot, individual study estimates are plotted against their corresponding precision values. A symmetrical funnel-shaped distribution generally indicates that study outcomes are evenly distributed around the pooled effect estimate. In the present analysis, the majority of studies cluster symmetrically around the central axis, suggesting that the synthesized results are unlikely to be strongly influenced by selective reporting. Nevertheless, minor asymmetry is observable among a small subset of studies represented in Figure 3, particularly those with smaller sample sizes or exploratory experimental designs. Such deviations are commonly observed in emerging research areas, where early studies may report larger effect sizes due to methodological variability or limited sample populations. For example, experimental research investigating microbial metabolic activity or enzyme production often involves small-scale laboratory optimization studies, which may produce variable outcomes depending on environmental conditions and culture parameters (Mefteh et al., 2019). Similarly, ecological studies investigating population differentiation and adaptation in agricultural pests may report variable effect sizes depending on geographic sampling and host plant associations (Wang et al., 2020).
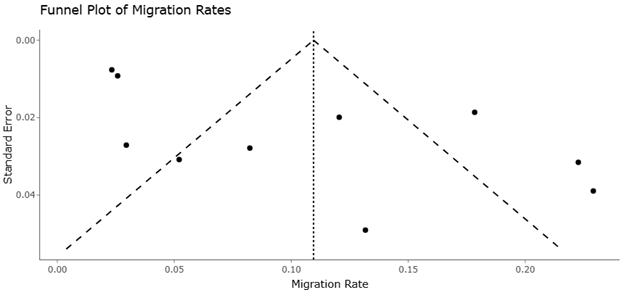
Figure 3. Funnel plot of Gene Flow Estimates for Sitobion avenae Biotypes. This plot displays migration rate estimates against their associated uncertainty to assess potential asymmetry or bias in gene flow estimates. The distribution provides a qualitative check on the robustness and consistency of Bayesian migration inferences across population contrasts.
Taken together, the forest and funnel plots provide an integrated statistical framework for interpreting the evidence synthesized in this review. The forest plot (Figure 2) demonstrates consistent biological responses across multiple studies while simultaneously revealing moderate heterogeneity in effect sizes. The funnel plot (Figure 3) indicates that the overall evidence base is relatively balanced, with only limited indications of potential bias. These complementary analyses strengthen the reliability of the synthesized findings and highlight the importance of integrating statistical visualization with biological interpretation when evaluating complex biological systems.