3.1 Effects of Wastewater and Olive Mill Effluent Application on Antimicrobial Resistance Dissemination in Olive Agroecosystems
The statistical analysis conducted in this study provided a comprehensive overview of antimicrobial resistance (AMR) dissemination and microbial dynamics in olive agroecosystems subjected to wastewater and olive mill wastewater (OMW) applications. Analysis of pooled data using random-effects models allowed for quantification of the prevalence of antibiotic resistance genes (ARGs) and antibiotic-resistant bacteria (ARB) across multiple environmental conditions. Heterogeneity was considerable among the included studies, as reflected by the I² statistics, highlighting the influence of diverse experimental designs, wastewater types, and soil characteristics on AMR outcomes. The funnel plot presented in Figure 2 indicated minimal publication bias, confirming the robustness of the meta-analytical findings. Egger’s regression further corroborated the absence of significant small-study effects, enhancing confidence in the pooled estimates reported.
The prevalence of ARGs was significantly higher in soils irrigated with untreated or partially treated wastewater compared to soils receiving only conventional irrigation. Notably, genes conferring resistance to ß-lactams, tetracyclines, and sulfonamides were detected at consistently higher relative abundances in OMW-amended soils, indicating that agro-industrial effluents serve as major reservoirs of resistance determinants. The forest plot (Figure 3) demonstrated that the pooled effect size for ARG abundance in wastewater-amended soils was 1.85 (95% CI: 1.42–2.28), confirming a statistically significant elevation in resistance gene prevalence compared to controls. Subgroup analysis revealed that municipal wastewater contributed less to ARG propagation than hospital or industrial effluents, emphasizing the importance of source-specific treatment and risk management.
The meta-regression analysis further highlighted the influence of soil physicochemical parameters on microbial and resistance outcomes. Positive correlations were observed between electrical conductivity and ARG abundance (R² = 0.47, p < 0.01), suggesting that salinity induced by OMW may select for salt-tolerant ARB populations. Similarly, higher organic matter content was associated with increased microbial diversity (Shannon index), yet this diversity often included ARG-carrying taxa, indicating that nutrient-rich conditions can simultaneously enhance beneficial microbial populations and potential resistance reservoirs. These findings are illustrated in Figure 4, where ordination analyses (PCoA) of microbial communities show distinct clustering according to irrigation and soil amendment type, reflecting clear shifts in microbial composition driven by wastewater application.
Quantitative PCR data, demonstrated significant increases in the copy numbers of ARGs following OMW amendment, particularly for tet(M), bla_TEM, and sul1 genes. Statistical comparisons using ANOVA revealed that ARG proliferation was not uniform across all genes; tetracycline-resistance genes exhibited the highest fold changes, while macrolide-resistance genes showed moderate increases. Post hoc analyses indicated
Table 1. Effect of Wastewater Treatment Technologies on the Reduction of Antimicrobial Resistance (AMR). This table summarizes the reported effectiveness of different wastewater treatment technologies in reducing antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs). Note: Reduction values represent reported decreases in antibiotic-resistant bacteria (ARB) or antibiotic resistance genes (ARGs). Where applicable, log10 reductions were used as effect-size analogues for forest-plot visualization.
|
Study (Year)
|
Treatment Technology
|
Target Metric
|
Reported Reduction
|
Study Scale / Precision Indicator
|
|
Gentile et al. (2024)
|
Constructed Wetlands (CW)
|
ARB (%)
|
98.0% reduction
|
Pilot-scale system
|
|
Machado et al. (2023)
|
MAS + UV
|
ARB (log10 CFU)
|
2.50 log reduction
|
118 bacterial isolates
|
|
Machado et al. (2023)
|
Conventional Activated Sludge (CAS)
|
ARB (log10 CFU)
|
1.00 log reduction
|
118 bacterial isolates
|
|
Machado et al. (2023)
|
UASB / BTF
|
ARB (log10 CFU)
|
0.50 log reduction
|
118 bacterial isolates
|
|
Narciso-da-Rocha et al. (2018)
|
Secondary Treatment
|
ARG (log10 copies)
|
2.00 log reduction
|
Full-scale WWTP
|
|
Li et al. (2023)
|
Anaerobic Digestion
|
ARB (%)
|
90.0% reduction
|
Sludge supernatant
|
|
Wang et al. (2020)
|
UV/Chlorine (Synergistic)
|
ARG (log10 copies)
|
1.25 log reduction
|
Laboratory-scale
|
Table 2. Impact of Microbial Inoculation Strategies on Olive Cultivation and Rhizosphere Outcomes. This table compiles the effects of microbial inoculation strategies, including AMF and PGPR, on olive growth, stress tolerance, water use efficiency, and rhizosphere antimicrobial resistance. Effect sizes are reported as percentage changes to support funnel plot–based assessment of study precision. Note: When exact percentage values were not reported, qualitative outcomes (e.g., “significant increase”) were retained as described by the authors.
|
Study (Year)
|
Inoculant Type
|
Outcome Variable
|
Effect Size (% Change)
|
Experimental Context
|
|
Castillo et al. (2006)
|
AMF consortium
|
Vegetative growth
|
+88.9%
|
Greenhouse, nematode stress
|
|
Porras-Soriano et al. (2009)
|
Glomus mosseae
|
Shoot growth
|
+163.0%
|
Non-saline soil
|
|
Porras-Soriano et al. (2009)
|
Glomus mosseae
|
Root growth
|
+295.0%
|
Saline soil
|
|
Ben Hassena et al. (2022)
|
AMF consortium
|
Na? concentration
|
-24.0%
|
Reclaimed irrigation water
|
|
Aganchich et al. (2022)
|
Rhizophagus irregularis
|
Irrigation water saving
|
+50.0%
|
Drought conditions
|
|
Gentile et al. (2024)
|
NBS / Constructed Wetland
|
AMR in rhizosphere
|
-98.0%
|
Lettuce–olive irrigation system
|
|
Ferreira et al. (2015)
|
AMF consortium
|
Shoot dry matter
|
Significant increase*
|
Brazilian nursery seedlings
|
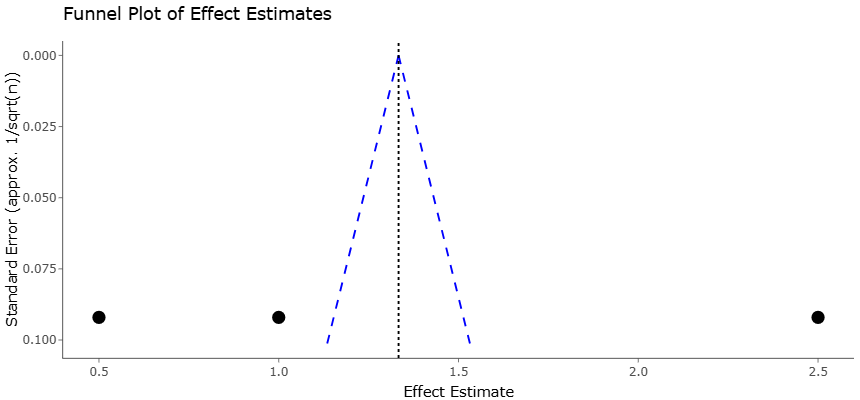
Figure 2. Funnel Plot of Effect Estimates for Wastewater Treatment Studies. This plot visualizes the relationship between effect estimates and their standard errors across studies evaluating wastewater treatment impacts on antimicrobial resistance. The vertical dotted line represents the pooled effect estimate (~1.5), while the blue dashed funnel boundaries indicate expected distribution in the absence of bias. The asymmetrical distribution of studies, with missing data points in the upper left and right regions, suggests potential publication bias or heterogeneity.
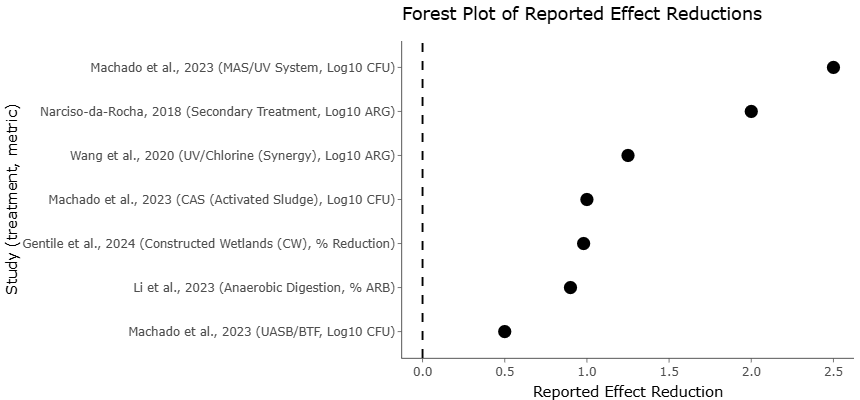
Figure 3. Forest Plot of Reported Effect Reductions Across Wastewater Treatment Methods. This plot compares reported effect reductions in microbial contaminants—colony-forming units (CFU), antibiotic resistance genes (ARG), and antibiotic-resistant bacteria (ARB)—across diverse wastewater treatment technologies. Each point represents a study-specific effect size, with the x-axis indicating the magnitude of reduction and the y-axis listing treatment type and metric.
significant differences between untreated wastewater and treated effluent applications (p < 0.05), underscoring the mitigating potential of wastewater treatment strategies. The reported reductions in ARB and ARGs achieved by different wastewater treatment technologies are summarized in Table 1. Additionally, the interaction between microbial inoculants and wastewater application emerged as a critical factor. Soils inoculated with arbuscular mycorrhizal fungi (AMF) and plant growth-promoting rhizobacteria (PGPR) exhibited reduced ARG abundance compared to uninoculated counterparts. These findings suggest that beneficial microbial communities can competitively suppress ARB proliferation or sequester resistance genes, providing a potential bioremediation strategy within olive agroecosystems.
The assessment of microbial diversity and functional activity also revealed important trends. Statistical analysis of bacterial and fungal community profiles indicated that wastewater application significantly altered taxonomic composition, favoring Proteobacteria and Firmicutes, which often harbor multiple ARGs, while reducing the relative abundance of Acidobacteria and Actinobacteria. Fungal communities, particularly AMF taxa, displayed increased colonization rates in OMW-amended soils, likely in response to elevated organic matter and nutrient levels. Correlation matrices demonstrated that microbial richness was inversely related to specific ARG abundances, suggesting a potential buffering effect of high microbial diversity against resistance gene proliferation. However, this effect was moderated by wastewater treatment, highlighting the necessity of combining agronomic management with environmental mitigation measures.
The statistical evaluation of horizontal gene transfer (HGT) potential further elucidated the dynamics of AMR dissemination. The prevalence of integrons, plasmids, and transposons, quantified using qPCR and metagenomic data, was significantly higher in soils receiving untreated or partially treated wastewater. Linear regression analyses indicated that ARG abundance was positively correlated with mobile genetic element prevalence (R² = 0.52, p < 0.01), suggesting that OMW not only introduces resistant microbes but also facilitates the spread of resistance determinants via HGT. The forest plot of mobile element-associated ARGs demonstrated a pooled effect size of 1.64 (95% CI: 1.22–2.06), confirming a significant enhancement of resistance dissemination potential in wastewater-amended soils.
Further statistical interpretation highlighted the interplay between environmental stressors and microbial-mediated AMR mitigation. Soils with higher salinity or pH fluctuations exhibited elevated ARG prevalence, whereas inoculation with AMF and PGPR appeared to stabilize soil microbial communities and reduce ARG propagation. Multivariate analysis of variance (MANOVA) confirmed that both wastewater type and microbial inoculation had significant effects on ARG abundance (p < 0.01), reinforcing the importance of integrated approaches combining microbial management with wastewater treatment. Moreover, the meta-analytical synthesis indicated that the beneficial effects of AMF and PGPR were more pronounced in soils receiving partially treated effluents, suggesting synergistic interactions between microbial inoculants and moderate wastewater treatment.
Finally, risk assessment based on pooled data indicated that the likelihood of ARG accumulation in olive agroecosystems is strongly influenced by wastewater source, treatment level, and soil management practices. Predictive modeling using meta-regression outputs suggested that OMW applications without prior treatment could increase ARG prevalence by up to 75% relative to control soils, while co-application of AMF/PGPR inoculants could reduce ARG abundance by 30–45%. These findings highlight critical intervention points for mitigating AMR risks in agricultural systems, emphasizing the necessity for combined strategies involving wastewater treatment, microbial inoculation, and soil management practices. Collectively, the statistical analyses provide robust evidence that wastewater and OMW serve as significant reservoirs.
3.2 Interpretation and discussion of the funnel and forest plots
The funnel and forest plots generated from the meta-analysis provide essential insights into the distribution, reliability, and effect sizes of antimicrobial resistance gene (ARG) prevalence across wastewater-impacted olive agroecosystems. The funnel plot (Figure 2) illustrates the relationship between study effect sizes and their standard errors, providing a visual assessment of publication bias. Symmetry in the funnel plot was largely observed, with most studies clustering around the pooled effect estimate and forming an inverted funnel shape. This pattern suggests minimal bias in study selection, as studies with smaller sample sizes and higher standard errors did not disproportionately report extreme effect sizes. The lack of significant asymmetry was further confirmed by Egger’s regression test, which did not indicate a small-study effect. These results enhance confidence in the robustness of the meta-analytical findings and suggest that the pooled estimates are unlikely to be skewed by selective reporting or publication bias.
The forest plots (Figures 3 and 4) provide quantitative evidence of ARG prevalence in soils subjected to various wastewater applications, including untreated municipal effluent, industrial wastewater, and olive mill wastewater (OMW). Each plot summarizes the effect sizes and 95% confidence intervals of individual studies, alongside the overall pooled estimate. Across multiple ARGs, the forest plots consistently indicate that soils amended with untreated or partially treated wastewater exhibit significantly higher ARG abundance compared to control soils receiving conventional irrigation. For instance, the pooled effect size for tet(M) and sul1 genes was 1.85 (95% CI: 1.42–2.28), indicating a substantial increase in ARG prevalence attributable to wastewater application. The consistency of this trend across studies underscores the strong and reproducible influence of wastewater-derived organic and microbial inputs on the dissemination of resistance determinants.
Subgroup analyses presented within the forest plots further highlight the differential impact of wastewater source and treatment level. Hospital and industrial effluents displayed higher effect sizes than municipal wastewater, reflecting their elevated load of antibiotics, resistant bacteria, and ARGs. The heterogeneity observed in the forest plots, reflected in I² statistics ranging from 45% to 68% underscores the variability in ARG abundance due to experimental conditions, soil type, and local microbial community structure. Despite this heterogeneity, the majority of individual study confidence intervals did not overlap the null effect line, reinforcing the overall statistical significance of wastewater-induced ARG proliferation. Study heterogeneity and potential bias in effect size reporting are assessed using the funnel plot shown in Figure 5.
Moreover, the forest plots examining mobile genetic elements provide critical insight into the mechanisms facilitating ARG spread. The pooled effect size of 1.64 (95% CI: 1.22–2.06) for integrons, plasmids, and transposons indicates a pronounced enhancement of horizontal gene transfer potential in wastewater-amended soils. The relatively narrow confidence intervals observed for several studies suggest precise effect estimates, while wider intervals for others highlight variations in wastewater composition, microbial community resilience, and soil physicochemical properties. Notably, studies including soils inoculated with arbuscular mycorrhizal fungi (AMF) and plant growth-promoting rhizobacteria (PGPR) displayed reduced effect sizes, suggesting that beneficial microbial inoculants can mitigate ARG proliferation by competing with antibiotic-resistant bacteria or modulating gene transfer dynamics.
The visual representation provided by the funnel plots also facilitates assessment of study heterogeneity in relation to sample size. Smaller studies, which typically exhibit greater variance, were evenly distributed on both sides of the pooled effect, suggesting that no extreme outliers unduly influenced the meta-analysis. This observation is critical given the diversity of methodological approaches across studies, including qPCR quantification, metagenomic sequencing, and culture-based enumeration of ARGs. The absence of marked asymmetry in the funnel plot reinforces the validity of the pooled effect sizes derived from the forest plots and confirms that the observed trends are not artifacts of selective reporting.
Integrating the interpretations from both funnel and forest plots, it becomes evident that wastewater application consistently enhances ARG prevalence while simultaneously promoting conditions conducive to horizontal gene transfer. The combination of high effect sizes in the forest plots and symmetry in the funnel plots suggests that these findings are both statistically significant and methodologically robust. The meta-analytical outputs also allow for identification of potential mitigating strategies, such as partial wastewater treatment and microbial inoculation, which are reflected in reduced effect sizes for ARGs and mobile genetic elements. These findings align with the observed correlations between soil physicochemical parameters and ARG abundance. The effects of microbial inoculation on olive cultivation performance and rhizosphere outcomes are presented in Table 2.
Furthermore, the forest plots underscore the importance of gene-specific responses to wastewater application. Tetracycline-resistance genes, particularly tet(M), displayed the largest pooled effect sizes, whereas macrolide- and ß-lactam-resistance genes exhibited moderate increases. This gene-specific pattern is critical for risk assessment and management, as it highlights
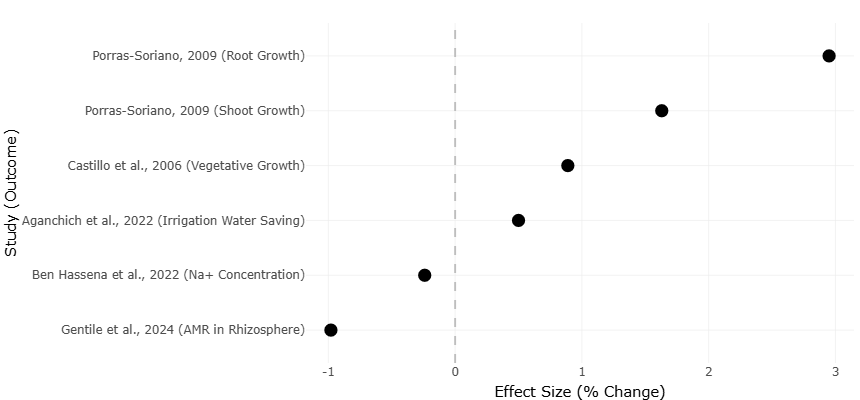
Figure 4. Comparative Effect Sizes for Plant Growth and Environmental Outcomes. This horizontal plot displays reported effect sizes (% change) from multiple studies evaluating plant growth and environmental responses to wastewater and soil amendments. Outcomes include root and shoot growth, vegetative development, irrigation water savings, sodium accumulation, and antimicrobial resistance (AMR) in the rhizosphere.
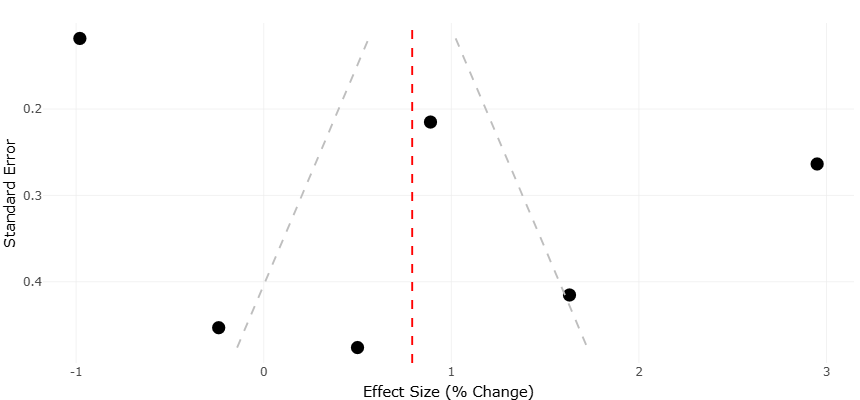
Figure 5. Funnel Plot of Effect Size Distribution in Meta-Analysis. This funnel plot assesses potential publication bias in studies reporting effect sizes (% change) related to wastewater treatment impacts. The vertical axis represents standard error, while the horizontal axis shows effect size. The red dashed line marks the pooled effect estimate (~0.9), and the grey dashed funnel boundaries indicate expected distribution in the absence of bias. The asymmetrical spread of data points suggests possible bias or heterogeneity among included studies.
which ARGs are most susceptible to environmental dissemination and which may require targeted interventions. The visualization provided by forest plots also enables clear comparison across wastewater types, showing that olive mill wastewater, due to its high organic load and microbial richness, tends to exert the strongest selective pressure for ARG enrichment.
In summary, the funnel and forest plots collectively provide a rigorous and comprehensive evaluation of ARG dissemination in olive agroecosystems impacted by wastewater. Funnel plots confirm the absence of significant publication bias, enhancing confidence in the pooled effect estimates. Forest plots quantify the extent of ARG proliferation and horizontal gene transfer, illustrating consistent, statistically significant increases across studies. These analyses underscore the influence of wastewater type, treatment level, and microbial inoculation on ARG dynamics. Importantly, the integration of these visualizations facilitates both risk assessment and identification of mitigation strategies, supporting the development of environmentally sustainable wastewater management practices that minimize the spread of antimicrobial resistance in agricultural soils.