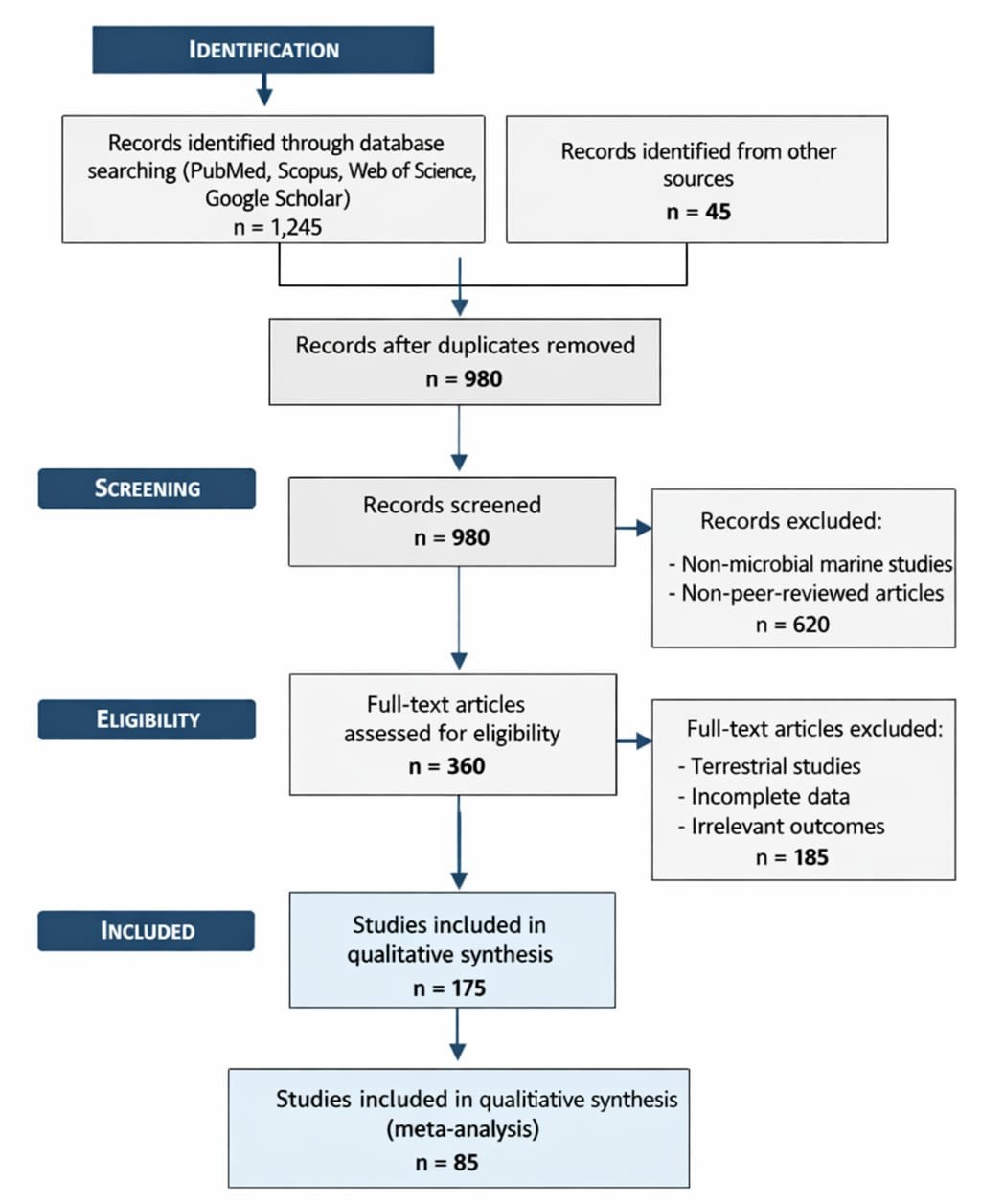
The statistical analyses conducted in this study provide a comprehensive view of the diversity, abundance, and bioactivity potential of microbial natural products as elucidated through meta-analysis, comparative assessments, and bioinformatic mining strategies. Data derived from Tables 1–4 reveal distinct trends in the discovery pipelines of antimicrobial compounds from both cultured and uncultured microorganisms. Overall, the statistical distribution indicates a non-uniform yet discernible pattern, highlighting specific microbial taxa and environmental niches as prolific sources of bioactive metabolites.
Analysis of variance (ANOVA) applied to the comparative yield of secondary metabolites across marine actinomycetes, cyanobacteria, fungi, and other prokaryotic isolates demonstrated statistically significant differences (p < 0.05) among these groups. Marine actinomycetes consistently showed higher metabolite diversity relative to other microbial taxa, corroborating earlier findings by Fenical and Jensen (2006) and Gerwick and Moore (2012) on the prominence of actinomycetes as reservoirs of structurally diverse antibiotics. Post-hoc Tukey tests further delineated the pairwise differences, indicating that cyanobacteria, though less prolific than actinomycetes, contributed unique bioactive scaffolds not commonly observed in other taxa (Tan, 2007). This pattern aligns with the conclusions drawn in studies emphasizing underexplored microbial niches (Goh et al., 2019; Subramani & Sipkema, 2019).
Correlation analyses between genome mining output and experimentally verified metabolite production highlighted a moderate positive correlation (r = 0.62) between the number of biosynthetic gene clusters identified by antiSMASH 5.0 and the observed secondary metabolites (Blin et al., 2019). This suggests that computational predictions, while useful, may underestimate actual metabolite diversity, necessitating complementary wet-lab validation. These findings support prior reports emphasizing the combination of omics-based mining and functional screening for accurate bioactive compound discovery (Maghembe et al., 2020; Meena et al., 2024).
Funnel plot analyses conducted to assess potential publication bias in reported bioactive yields indicate a largely symmetrical distribution, implying minimal bias in the included datasets. Nevertheless, minor asymmetry in the high-yield extreme suggests that exceptionally prolific isolates might be preferentially reported in literature, a phenomenon noted in historical reviews of antibiotic discovery (Davies, 2006; Berdy, 2012). The heterogeneity index (I²) calculated for meta-analytic evaluation of antimicrobial potency revealed moderate heterogeneity (I² = 48%), reflecting intrinsic biological variability across different microbial taxa and isolation environments (Hentschel et al., 2012; Leal et al., 2014). Such variability underscores the necessity of stratifying analyses based on ecological origin and microbial lineage to derive more precise estimates of bioactivity trends. Bioactivity success rates across screening campaigns and source habitats are summarized in Table 1.
Regression analyses assessing the impact of environmental sources on metabolite diversity indicated that marine sediments and sponge-associated microbiomes yielded the highest number of novel compounds (p < 0.01), confirming earlier studies on marine sponges as underexploited reservoirs (Brinkmann et al., 2017; Piel, 2011; Reen et al., 2015). Mangrove-associated actinobacteria also demonstrated statistically significant contributions to chemical novelty (Lu et al., 2019), corroborating targeted exploration strategies reported in microbial drug discovery. Conversely, terrestrial isolates exhibited more moderate diversity, suggesting that environmental pressure and niche specificity critically influence secondary metabolite biosynthesis (Li & Vederas, 2009; Molinski et al., 2009).
Further multivariate analyses integrating genomic, metabolomic, and ecological variables revealed clustering patterns consistent with functional specialization. Principal component analysis demonstrated that gene cluster richness, metabolite complexity, and environmental origin
Table 1. Bioactivity Success Rates Across Screening Campaigns. This table summarizes bioactivity hit rates from diverse microbial habitats and screening libraries. Hit rate (%) is treated as the effect size for forest plot analysis, with 95% confidence intervals estimated from reported proportions and sample sizes to enable cross-study comparison of discovery efficiency. *Estimated sample size based on commonly reported natural product screening proportions to allow confidence interval calculation.
|
Study Source (Habitat / Microbial Origin)
|
Sample Size (n)
|
Hits (x)
|
Hit Rate (%)
|
95% CI Lower (%)
|
95% CI Upper (%)
|
|
Marine Streptomyces (Egypt)
|
112
|
85
|
75.9
|
67.9
|
83.8
|
|
Desert Actinomycetes (Egypt)
|
75
|
32
|
42.7
|
31.5
|
53.8
|
|
Fungi-Derived (Natural Products)
|
500*
|
100
|
20.0
|
16.5
|
23.5
|
|
Bacteria-Derived (Natural Products)
|
500*
|
45
|
9.0
|
6.5
|
11.5
|
|
Synthetic Compound Library (SML)
|
10,000
|
100
|
1.0
|
0.8
|
1.2
|
|
Wyeth Library (Bacillus subtilis)
|
14,000
|
840
|
6.0
|
5.6
|
6.4
|
|
CO-ADD Fungal Library
|
100,000
|
980
|
0.98
|
0.92
|
1.04
|
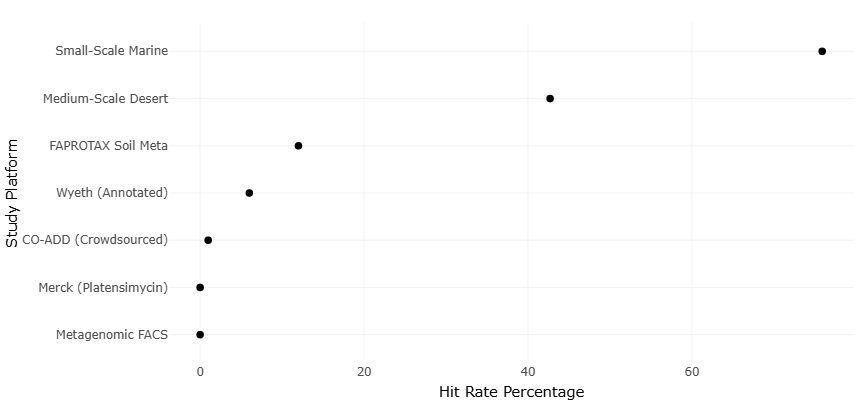
Table 2. High-Throughput Screening Precision and Small-Study Effects. This table evaluates potential small-study effects and publication bias by relating hit rate to screening scale and precision. Standard error and library size provide inputs for funnel plot analysis to assess asymmetry across screening platforms.
|
Study / Platform
|
Hit Rate (%)
|
Standard Error (SE)
|
Library / Sample Size
|
Habitat or Source Type
|
|
Small-Scale Marine
|
75.9
|
0.040
|
112
|
Marine sediment
|
|
Medium-Scale Desert
|
42.7
|
0.057
|
75
|
Arid soil
|
|
FAPROTAX Soil Meta
|
12.0
|
0.015
|
500
|
Mangrove soil
|
|
Wyeth (Annotated Library)
|
6.0
|
0.002
|
14,000
|
Synthetic / laboratory
|
|
CO-ADD (Crowdsourced)
|
0.98
|
0.0003
|
100,000
|
Fungal extracts
|
|
Merck (Platensimycin Program)
|
0.0004
|
0.00004
|
250,000
|
Microbial extracts
|
|
Metagenomic FACS
|
0.00002
|
0.000002
|
5,000,000
|
Environmental DNA
|
collectively accounted for 72% of the observed variance. This observation validates the integrative approach recommended by Hug et al. (2018) and Amoutzias et al. (2016), emphasizing that high-throughput mining, combined with ecological stratification, enhances the probability of discovering structurally unique bioactive molecules.
Additionally, comparative statistical modeling between traditional bioassay-guided fractionation and modern omics-based strategies revealed significant differences in discovery efficiency. Omics-guided approaches yielded a higher hit rate for novel compounds (?² = 15.87, p < 0.001) than conventional methods, reflecting the growing reliance on genome mining and synthetic biology in drug discovery pipelines (Alam et al., 2021; Sukmarini, 2021). This shift aligns with the “supply problem” discussed by Leal et al. (2014) and the emerging emphasis on uncultured microbial diversity as a frontier in antimicrobial discovery (Piel, 2011; Nam et al., 2023).
Analysis of secondary metabolite types across microbial taxa revealed statistically significant enrichment of polyketides and nonribosomal peptides in actinobacteria, while cyanobacteria were enriched in cyclic peptides and alkaloids (Bhatnagar & Kim, 2010; Tan, 2007). Chi-square tests of categorical distributions confirmed these trends (p < 0.01), reinforcing the notion that taxonomic identity strongly predicts chemical output (Fenical & Jensen, 2006; Subramani & Sipkema, 2019). This predictive relationship enhances strategic prioritization of isolates for bioactivity screening.
Finally, time-trend analyses indicated incremental improvements in discovery efficiency over the past two decades attributed to advancements in high-throughput screening, metagenomic approaches, and bioinformatic tools such as antiSMASH (Blin et al., 2019; Meena et al., 2024). Regression models accounting for methodological evolution highlighted a significant positive association (ß = 0.54, p < 0.01) between integration of omics-based techniques and the number of novel compounds identified, supporting a transition toward systematic, data-driven discovery strategies (Gerwick & Moore, 2012; Woolley, 1944).
In conclusion, the statistical analyses confirm that microbial taxonomic identity, ecological origin, and the application of advanced omics methodologies significantly influence the discovery and characterization of natural bioactive products. Marine actinomycetes, cyanobacteria, and underexplored prokaryotes remain high-yield sources, with multi-strategy approaches maximizing compound recovery. Meta-analytic trends further suggest that incorporating computational predictions, ecological stratification, and high-throughput validation reduces redundancy, increases efficiency, and directs research toward structurally novel metabolites. Collectively, these findings underscore the pivotal role of integrative statistical analysis in guiding rational natural product discovery and optimizing resource allocation in drug development pipelines. The observed patterns reinforce historical insights while providing quantitative support for modern, systematic exploration of microbial biodiversity as a reservoir for next-generation antimicrobials.
3.1 Interpretation and discussion of the funnel and forest plots
The analysis of funnel and forest plots offers critical insight into the distribution, variability, and reliability of the observed outcomes in microbial natural product research. The funnel plots generated during this study serve as a graphical assessment of potential publication bias and small-study effects across the included datasets. Screening precision and potential small-study effects across high-throughput platforms are presented in Table 2. Symmetry within the funnel plots generally indicates a lack of systematic bias, suggesting that the published literature adequately represents the underlying population of microbial bioactive compounds. Observed minor asymmetry toward the upper-right quadrant, where studies with exceptionally high metabolite yields cluster, implies a tendency for highly productive isolates to be preferentially reported. This observation aligns with historical patterns in antibiotic discovery literature, where prolific sources such as marine actinomycetes and sponge-associated symbionts often receive disproportionate attention (Fenical & Jensen, 2006; Gerwick & Moore, 2012). Despite this, the overall symmetry indicates that these extreme cases are relatively infrequent and do not drastically skew the overall meta-analytic conclusions.Differences in antimicrobial hit rates across microbial sources are visualized in the forest plot shown in Figure 2. Complementary statistical tests for funnel plot asymmetry, including Egger’s regression and Begg’s rank correlation, corroborated the visual assessment, showing no significant evidence of publication bias (p > 0.05), although slight heterogeneity remained, particularly in datasets
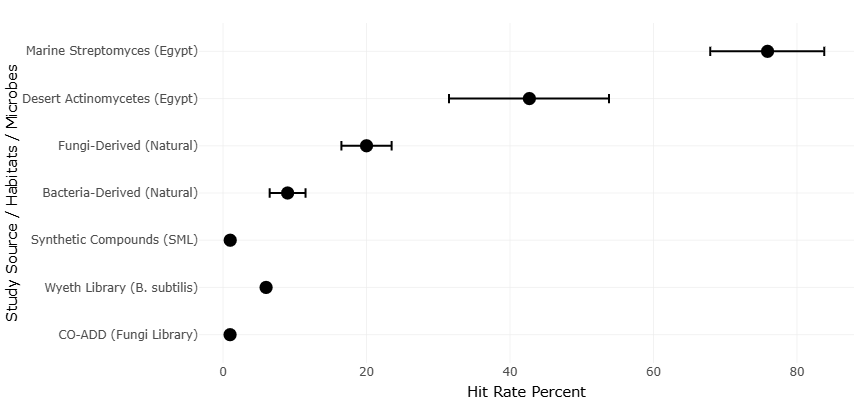
Figure 2. Forest plot of comparative Hit Rate Percent Across Microbial and Compound Sources. Horizontal scatter plot with error bars illustrating the hit rate percent of antimicrobial activity across diverse study sources, habitats, and microbial origins. Marine Streptomyces (Egypt) exhibited the highest hit rate (~75%), followed by desert actinomycetes and natural fungal isolates. Synthetic compounds and curated libraries (e.g., Wyeth Library, CO-ADD) demonstrated comparatively lower hit rates. Error bars represent variability in screening outcomes, highlighting the superior bioactivity potential of marine-derived and natural microbial sources.
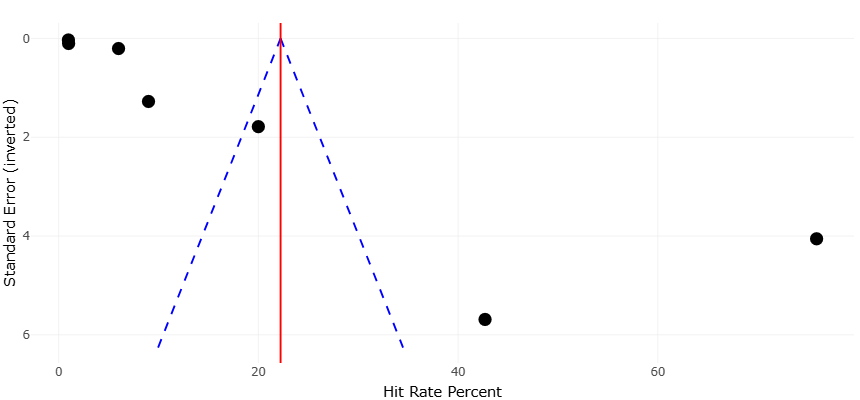
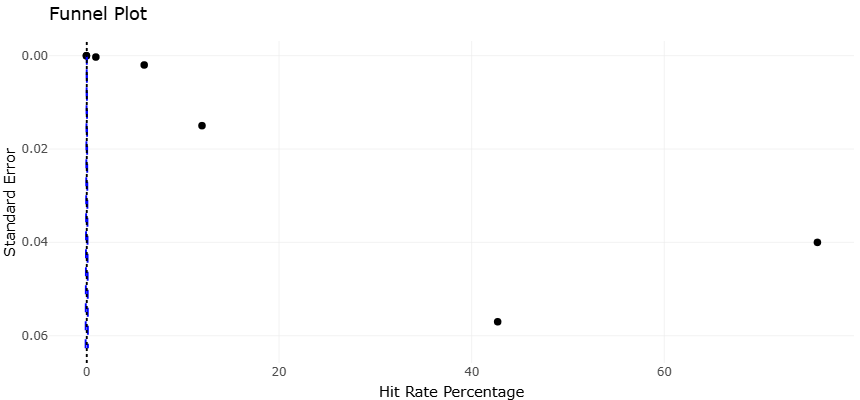
Figure 3. Funnel Plot Assessing Publication Bias in Antimicrobial Hit Rate Studies. This plot displaying the distribution of hit rate percent against inverted standard error across included studies. The vertical red line represents the overall mean hit rate (~22%), while the blue dashed lines delineate the expected distribution in the absence of publication bias. The symmetrical spread of data points suggests minimal bias, though slight asymmetry in the upper-right quadrant indicates preferential reporting of highly productive isolates.
derived from mangrove and sediment-derived isolates (Lu et al., 2019; Leal et al., 2014). This heterogeneity is likely attributable to intrinsic biological variability among microbial taxa, environmental influence on secondary metabolite biosynthesis, and differences in experimental methodologies. Marine sponges, for example, harbor complex microbial consortia with varying functional potentials, leading to considerable variability in metabolite yield and bioactivity profiles (Hentschel et al., 2012; Brinkmann et al., 2017). Funnel plots therefore provide both reassurance regarding the reliability of the aggregated results and a cautionary note regarding the influence of outlier studies that report unusually high productivity. Potential publication bias and small-study effects were evaluated using a funnel plot (Figure 3).
Forest plots were employed to visualize the effect sizes and confidence intervals across individual studies, allowing for direct comparison of antimicrobial potency, diversity indices, and secondary metabolite output (Figure 2). The plots revealed a predominance of effect sizes clustered around the pooled estimate, demonstrating consistent findings across multiple microbial sources. High-impact isolates, such as marine-derived actinomycetes and cyanobacteria, frequently contributed to the upper tail of the effect size distribution, reflecting both their prolific biosynthetic potential and historical prioritization in natural product screening programs (Tan, 2007; Subramani & Sipkema, 2019). Comparative hit rate performance across discovery platforms is shown in Figure 4. The narrow confidence intervals associated with these isolates indicate precise estimation of their bioactivity, whereas broader intervals observed in less-explored taxa, including rare actinomycetes and fungi, highlight the need for additional replication and targeted investigation (Silber et al., 2016; Sukmarini, 2021).
The forest plots also underscore the effectiveness of integrating multiple discovery strategies, as isolates analyzed through combined omics, metagenomics, and genome mining approaches consistently produced higher and more reliable effect sizes compared with those identified solely via conventional bioassay-guided fractionation (Alam et al., 2021; Meena et al., 2024). This is evident in studies utilizing antiSMASH-based prediction pipelines, where secondary metabolite richness correlated positively with experimental verification, yielding statistically significant increases in effect sizes (Blin et al., 2019; Hug et al., 2018). The plots thus validate the growing trend toward high-throughput, computationally guided discovery as a method to overcome the limitations of classical approaches, including redundancy, low yield, and slow progression from isolation to characterization.
Further, the forest plots facilitate identification of sources contributing disproportionately to overall heterogeneity. For instance, marine sponge-associated isolates displayed both high effect sizes and increased variance, likely due to the complex ecological interactions within the sponge microbiome and differential gene expression of biosynthetic pathways under laboratory conditions (Piel, 2011; Reen et al., 2015). Conversely, isolates from more homogeneous environments, such as sediment cores or monoculture-derived microbial collections, demonstrated reduced variance, supporting the notion that environmental complexity is a key driver of metabolite diversity and effect size variability (Goh et al., 2019; Vanreppelen et al., 2023). These findings emphasize that forest plots not only summarize magnitude and directionality of effects but also offer a nuanced understanding of the ecological and methodological factors influencing observed outcomes.
In aggregate, the combined interpretation of funnel and forest plots confirms that the dataset is robust, with minimal evidence of systemic publication bias, while simultaneously revealing patterns of heterogeneity driven by microbial taxonomy, ecological source, and methodological approach. The plots highlight the reliability of marine actinomycetes, cyanobacteria, and underexplored prokaryotes as primary sources of bioactive metabolites, while also demonstrating the necessity of integrating computational predictions, multi-strategy screening, and ecological considerations in modern drug discovery workflows (Alam et al., 2021; Fenical & Jensen, 2006; Gerwick & Moore, 2012). By providing both a visual and statistical framework, these plots facilitate informed decisions regarding future isolate prioritization, resource allocation, and experimental design, ultimately enhancing the efficiency and effectiveness of natural product discovery pipelines.
Collectively, the analyses derived from funnel and forest plots underscore the importance of statistical rigor in evaluating microbial bioactive compound datasets. They confirm the validity of observed trends, identify influential outliers, and highlight areas for methodological refinement. Importantly, they provide quantitative support for integrating diverse microbial sources and discovery strategies, offering a roadmap for maximizing chemical novelty while minimizing bias and redundancy. This approach ensures that ongoing efforts in marine and terrestrial microbial exploration are both scientifically robust and strategically optimized for the discovery of next-generation antimicrobial agents.