1. Introduction
Antimicrobial resistance (AMR) is no longer a distant or abstract concern—it is, increasingly, a defining feature of modern medicine’s limitations. What once appeared controllable has, over time, revealed a far more complex and persistent character. Treatments that were once routine now carry uncertainty, and the broader implications are difficult to ignore. There is a growing sense that the medical community is not merely facing isolated failures of antibiotics, but rather a systemic erosion of their reliability. Estimates suggesting millions of annual deaths in the coming decades, alongside profound economic consequences, underscore the scale of this challenge (O’Neill, 2016). Yet, even beyond such projections, the day-to-day realities—treatment failures, prolonged infections, and rising healthcare burdens—signal that AMR is already deeply embedded within global health systems.
Historically, the story of antibiotics has been one of remarkable discovery followed, somewhat unexpectedly, by stagnation. The mid-twentieth century marked an era of extraordinary productivity, often referred to as the “Golden Age” of antibiotic discovery, when soil-derived microorganisms—particularly actinomycetes—yielded a wealth of clinically transformative compounds (Berdy, 2012; Colegate & Molyneux, 2008). However, this momentum gradually slowed. By the late twentieth century, the repeated rediscovery of known molecules, combined with declining industrial incentives, created what many now describe as a “discovery void” (Levy & Marshall, 2004). The pipeline, once abundant, became increasingly sparse, raising uncomfortable questions about whether the limits of conventional discovery strategies had already been reached.
At the same time, our understanding of resistance itself has undergone a subtle but important shift. It is now clear that antibiotic resistance is not simply a byproduct of modern clinical misuse; rather, it is deeply rooted in microbial evolution. Evidence demonstrating the ancient origin of resistance genes suggests that these traits predate human antibiotic use by millions of years (D’Costa et al., 2011). This realization complicates the narrative. Resistance is not an anomaly—it is, in many ways, a natural and expected outcome of microbial ecology. Consequently, AMR must be understood not only as a clinical issue but also as an ecological and evolutionary phenomenon shaped by interactions across diverse environments (Martinez, 2009 Stephen et al., 2008).
This broader ecological perspective has, perhaps inevitably, redirected attention toward nature itself—not as a static repository of known compounds, but as a dynamic and largely unexplored source of chemical diversity. Environments once considered peripheral to drug discovery—deep-sea sediments, extreme habitats, and symbiotic microbial niches—are now recognized as potential reservoirs of novel bioactive molecules (Bull & Stach, 2007). These settings often impose unique selective pressures, encouraging microorganisms to evolve specialized metabolic pathways that may produce structurally and functionally distinct compounds.
Marine ecosystems, for instance, illustrate this complexity in striking ways. Microbial communities associated with sponges, plastics, and phytoplankton create microenvironments characterized by intense competition and chemical signaling (Grossart & Rojas-Jimenez, 2016; Keswani et al., 2016; Hentschel et al., 2012). Such interactions are not merely ecological curiosities—they can directly influence the production of secondary metabolites with antimicrobial properties. Similarly, host-associated microbiomes, including those found in insects, have revealed intricate symbiotic relationships in which microorganisms contribute to host defense through antibiotic production (Currie et al., 1999; Chevrette et al., 2019). These observations suggest that, rather than searching broadly and indiscriminately, there may be value in focusing on ecological contexts where chemical interactions are already highly refined.
Despite these promising directions, a persistent challenge remains: the vast majority of microorganisms cannot be readily cultivated using traditional laboratory techniques. It is estimated that only a small fraction of microbial diversity is accessible through standard methods, leaving an immense reservoir of biosynthetic potential unexplored (Lewis et al., 2010). This limitation has, in recent years, prompted a shift toward genome-based discovery approaches. Advances in sequencing technologies and bioinformatics now allow researchers to identify biosynthetic gene clusters (BGCs) directly from environmental DNA, offering insights into metabolic capabilities without the need for cultivation (Medema et al., 2011; Blin et al., 2021).
Yet, even this approach introduces new complexities. The presence of a gene cluster does not guarantee its expression. Many biosynthetic pathways remain silent under laboratory conditions, requiring specific environmental cues or interspecies interactions to become active. To address this, researchers have developed strategies such as co-culture systems and the One Strain, Many Compounds (OSMAC) approach, which aim to mimic natural ecological conditions and thereby stimulate the production of otherwise cryptic metabolites (Bode et al., 2002; Bertrand et al., 2014; Netzker et al., 2015; Gross et al., 2007). These methods, while promising, highlight the intricate relationship between microbial behavior and environmental context—a relationship that is not easily replicated in controlled settings.
In parallel, technological innovations have begun to reshape the scale and efficiency of antimicrobial discovery. Microfluidic platforms, for example, enable the high-throughput screening of microbial populations at the single-cell level, dramatically increasing the likelihood of identifying rare or low-abundance producers (Agresti et al., 2010). Such approaches represent a departure from traditional screening methods, offering a level of resolution and throughput that was previously unattainable. At the same time, advances in analytical chemistry, including mass spectrometry-based molecular networking, have improved the ability to distinguish novel compounds from known ones, reducing redundancy in the discovery process.
Still, despite these advances, the field remains fragmented. Different studies employ varying methodologies, target diverse ecological niches, and report outcomes using inconsistent metrics. This heterogeneity complicates efforts to draw broader conclusions about where and how antimicrobial discovery is most effective. While individual studies often highlight promising findings, there is a noticeable lack of synthesis across the field—a gap that becomes increasingly significant as research activity accelerates.
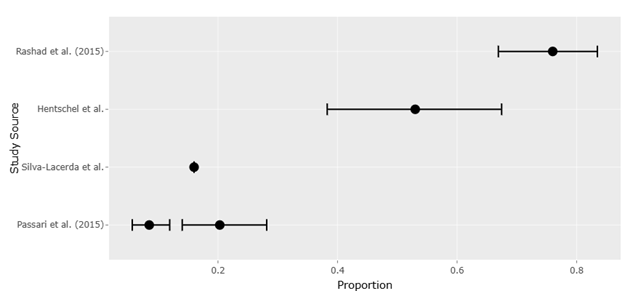
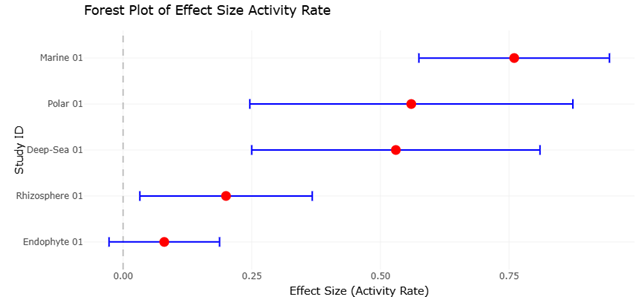
Against this backdrop, systematic reviews and meta-analytical approaches offer a way to bring coherence to an otherwise dispersed body of knowledge. By integrating findings across multiple studies, it becomes possible to identify patterns that are not immediately apparent at the individual study level. For instance, emerging evidence suggests that microorganisms derived from extreme or symbiotic environments may exhibit higher antimicrobial activity compared to those from more conventional sources, although such conclusions remain tentative due to methodological variability.
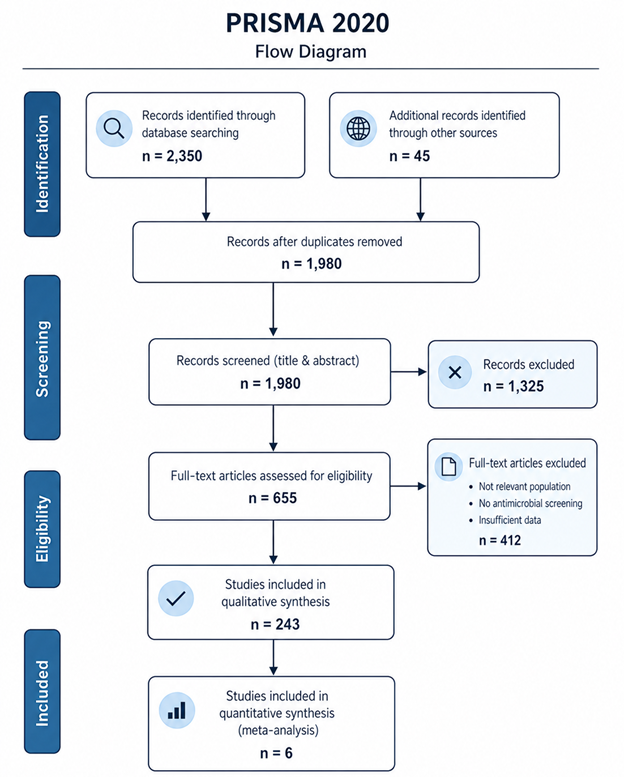
The present systematic review is situated within this evolving landscape. Rather than focusing on a single environment or methodological approach, it seeks to examine antibiotic bioprospecting from a broader, integrative perspective. By synthesizing evidence across ecological contexts, discovery strategies, and analytical frameworks, this study aims to clarify where meaningful progress is being made—and where limitations persist. In doing so, it hopes to contribute not only to the scientific understanding of antimicrobial discovery but also to the ongoing effort to address one of the most pressing challenges of our time.