1. Introduction
The microbial world—vast, intricate, and still only partially understood—extends across nearly every conceivable ecological boundary. From sunlit coastal waters to the immense pressures of the deep ocean, and from fertile soils to nutrient-starved cave systems, microorganisms persist in conditions that, at times, seem almost incompatible with life. It is perhaps this very adaptability that has drawn sustained scientific attention in recent decades. Increasingly, marine and pristine ecosystems are being recognized not merely as reservoirs of biodiversity, but as dynamic biochemical landscapes with tangible implications for biotechnology, medicine, and environmental sustainability.
What makes these environments particularly compelling is not just their diversity, but the selective pressures they impose. Extreme temperature gradients, salinity fluctuations, nutrient limitations, and hydrostatic pressure appear to drive microorganisms toward unusual metabolic solutions. These solutions often manifest as structurally complex secondary metabolites and specialized enzymes—sometimes referred to as extremozymes—that are rarely encountered in more conventional, terrestrial settings (Miller et al., 2017). In many ways, these molecules reflect evolutionary problem-solving at its most refined, shaped by constraints that demand efficiency and resilience.
At the same time, our capacity to study such systems has shifted quite dramatically. Earlier work in microbiology relied heavily on cultivation-based approaches, which, as is now widely acknowledged, capture only a fraction of existing microbial diversity. The so-called “great plate count anomaly” continues to remind us that most microorganisms remain uncultured and, therefore, historically overlooked (Daniel, 2005). The transition toward metagenomics and genome-resolved analyses has begun to address this gap, enabling researchers to access previously hidden lineages and metabolic pathways. Tools such as antiSMASH have further refined this process, allowing for systematic identification of biosynthetic gene clusters and providing insight into the chemical potential encoded within microbial genomes (Medema et al., 2011).
Marine systems, in particular, have emerged as especially rich in bioactive potential. It is now fairly well established that many compounds originally attributed to macroorganisms—sponges, corals, and tunicates—are, in fact, produced by associated microbial symbionts. Investigations into sponge-associated bacteria, for instance, have revealed an abundance of secondary metabolism gene clusters, particularly within Streptomyces species, that are capable of synthesizing compounds with potent antimicrobial activity (Jackson et al., 2018; Kennedy et al., 2009). These findings have contributed to a broader shift in perspective, one that views marine microbiomes not as passive inhabitants but as active biochemical contributors within their host systems.
Some of the most intriguing discoveries, however, have come from environments that are physically and chemically extreme. Deep-sea ecosystems, including the Mariana Trench, have yielded organisms such as Micromonospora provocatoris, which produce rare siderophores and macrolides with notable bioactivity (Abdel-Mageed et al., 2021). Similarly, hydrothermal vent systems support chemosynthetic microbial communities capable of generating enzymes that remain stable under high temperature and pressure—properties that are particularly attractive for industrial applications. These findings align with broader efforts in blue biotechnology, where marine-derived compounds are increasingly explored for pharmaceutical and industrial use (Maldonado-Ruiz et al., 2024; Labes, 2023).
Yet, it would be an oversimplification to assume that bioactive potential is confined to marine environments alone. Pristine terrestrial systems—especially those that are geographically isolated or resource-limited—offer equally compelling opportunities for discovery. Cave ecosystems, for example, present oligotrophic conditions that favor microorganisms with highly specialized metabolic capabilities. Actinobacteria isolated from such environments have been shown to produce compounds like cervimycins, which retain activity against antibiotic-resistant pathogens (Herold et al., 2005; Gatinho et al., 2023). In a somewhat different context, the cryosphere—encompassing glaciers and permafrost—hosts cold-adapted microbes that synthesize antifreeze proteins and extracellular polymeric substances, enabling survival under subzero conditions (Margesin & Collins, 2019; Beniston et al., 2018). These adaptations, while ecologically driven, also carry clear translational potential.
Beyond environmental extremity, microbial bioactivity is also evident in more unexpected settings. Cultural heritage sites, for instance, have been found to harbor microbial communities capable of both degrading and synthesizing complex organic compounds. While such activity is often discussed in the context of biodeterioration, it also highlights an underexplored biochemical capacity within anthropogenic environments. This broader ecological perspective—one that spans natural and human-influenced systems—suggests that microbial innovation is not confined to a single niche but is instead distributed across a continuum of habitats.
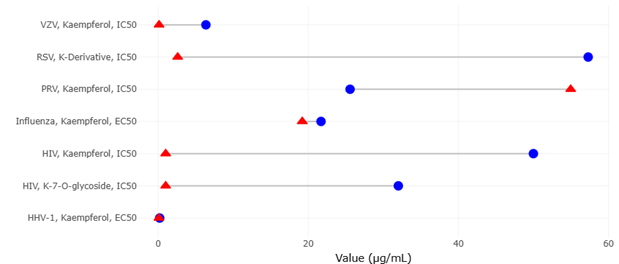
Among the various classes of microbial metabolites, flavonoids and related compounds have received particular attention due to their antiviral properties. Kaempferol, for instance, has been shown to exhibit activity against a range of viruses, including herpesviruses, HIV-1, and influenza, often through modulation of host signaling pathways (Behbahani et al., 2013; Behbahani et al., 2014). Subsequent studies have expanded on these findings, demonstrating inhibitory effects against varicella-zoster virus and pseudorabies virus, as well as broader antiviral potential across multiple systems (Park et al., 2022; Chen et al., 2021). Additional work has highlighted the role of flavonoid glycosides in antiviral activity, reinforcing the importance of structural diversity within this class of compounds (Zhou et al., 2014; Kai et al., 2014).
What is perhaps most striking is the consistency with which such compounds appear across diverse microbial sources. Reviews synthesizing this body of evidence suggest that kaempferol and related molecules represent a promising, though still incompletely understood, category of antiviral agents (Periferakis et al., 2023). At the same time, marine natural products more broadly continue to demonstrate an extraordinary range of bioactivities, from antimicrobial to neuroprotective effects, underscoring their relevance for drug discovery (Blunt et al., 2011). Compounds such as manzamine alkaloids, for example, have shown inhibitory activity against key enzymes implicated in neurodegenerative disease, hinting at applications beyond infectious disease (Hamann et al., 2007).
In parallel with therapeutic exploration, there has been growing interest in the application of microbial metabolites within agriculture. Plant growth-promoting microorganisms (PGPMs), when effectively delivered through encapsulation technologies, have been shown to enhance crop resilience and productivity under stress conditions (Balla et al., 2022; John et al., 2011). These approaches, often framed within the broader context of sustainable agriculture, reflect an effort to reduce reliance on chemical inputs while maintaining agricultural efficiency. Bioformulation strategies continue to evolve, integrating insights from microbiology, materials science, and agronomy (Ijaz et al., 2019).
Despite these advances, challenges remain. The interpretation of microbial genomic data, particularly in the context of biosynthetic potential, is not always straightforward. Similarities in sequence or structure do not necessarily translate to functional equivalence, as highlighted in recent comparative analyses of viral protein domains (Dixson et al., 2024). Such findings serve as a reminder that bioinformatic predictions must be approached with caution and, where possible, validated experimentally.
Taken together, the current body of research points toward a field that is both rapidly expanding and inherently interdisciplinary. Marine and pristine microbial ecosystems offer a wealth of chemical diversity, much of which remains untapped. At the same time, advances in sequencing, computational biology, and analytical chemistry are making it increasingly feasible to explore this diversity in a systematic and reproducible manner.
This narrative review, therefore, seeks to synthesize existing knowledge on microbial diversity and bioactive potential across marine and pristine environments. Rather than focusing solely on individual discoveries, it aims to situate these findings within a broader ecological and biotechnological framework. In doing so, it becomes possible—if only tentatively—to trace emerging patterns in microbial adaptation, metabolic innovation, and applied potential. Ultimately, such an integrative perspective may help guide future efforts in bioprospecting, ensuring that the exploration of microbial “chemical space” proceeds with both scientific rigor and practical relevance.