1. Introduction
The marine environment—vast, layered, and still only partially explored—has increasingly come to be seen as a reservoir of biochemical innovation. While it is well known that oceans cover the majority of the Earth’s surface, what remains less fully appreciated is the extent to which microbial life within these systems contributes to chemical diversity. Marine microorganisms, including bacteria, fungi, and cyanobacteria, inhabit ecological niches defined by fluctuating salinity, pressure, and nutrient availability, and these conditions appear to foster the evolution of highly specialized metabolic capabilities (Andryukov et al., 2019).
Within this context, secondary metabolites emerge as particularly intriguing. Unlike primary metabolites, which are essential for cellular survival, secondary metabolites are often associated with ecological interactions—competition, signaling, and defense. Yet the boundary between these categories is not always clear-cut, and in some cases, secondary metabolites exert effects that are central to survival under stress conditions (Arnison et al., 2013). Fungal systems, for example, demonstrate how metabolite production can be tightly linked to pathogenicity and environmental adaptation (Alves et al., 2025; Raffa & Keller, 2019).
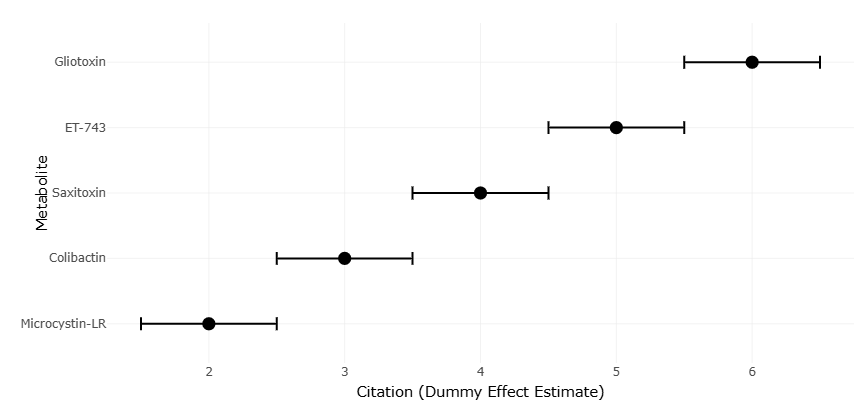
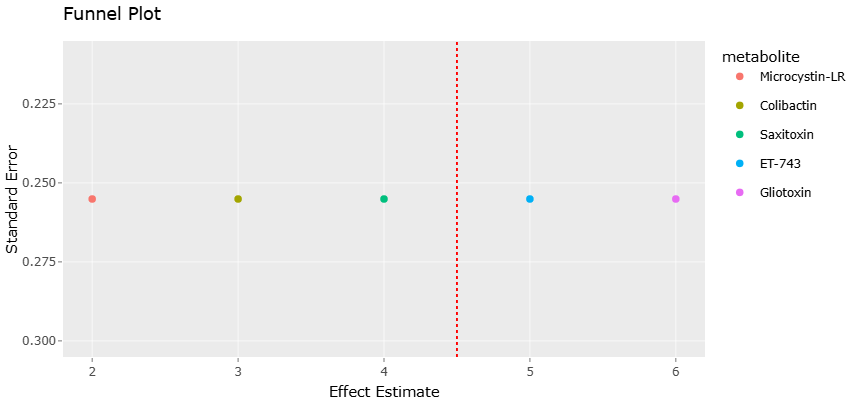
The growing urgency of antimicrobial resistance has, perhaps unsurprisingly, renewed interest in these compounds. Marine-derived metabolites have been shown to exhibit antibacterial, anticancer, and immunosuppressive activities, often with structural features that differ significantly from those found in terrestrial organisms (Amoutzias et al., 2016; Yamashita et al., 2015). Compounds such as ET-743 and bryostatins illustrate how marine symbioses can yield clinically relevant molecules, although their discovery often raises questions about the true microbial producers (Rath et al., 2011; Davidson et al., 2001).
At the same time, the dual nature of these metabolites cannot be ignored. Cyanobacteria, for instance, produce potent toxins such as microcystins and saxitoxins, which have significant ecological and public health implications (Tillett et al., 2000; Mihali et al., 2009). Similarly, bacterial genotoxins such as colibactin are capable of inducing DNA damage and contributing to disease processes, highlighting the fine line between therapeutic potential and toxicity (Nougayrède et al., 2006; Xue et al., 2019; Wilson et al., 2019).
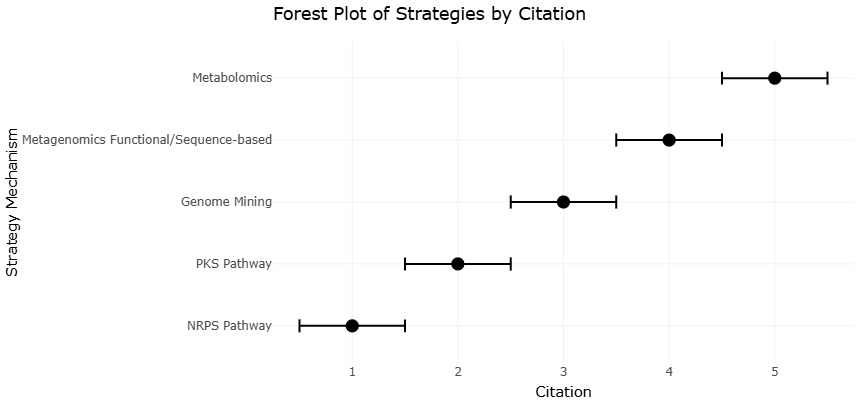
The remarkable diversity of these compounds is underpinned by sophisticated biosynthetic systems, particularly non-ribosomal peptide synthetases (NRPS) and polyketide synthases (PKS). These modular enzyme complexes function as assembly lines, incorporating diverse building blocks into structurally complex molecules (Strieker et al., 2010). Structural analyses have revealed that NRPS enzymes can adopt multiple conformations, which may contribute to their catalytic flexibility (Drake et al., 2016).
Polyketide biosynthesis follows a similarly modular logic, involving iterative condensation reactions and domain-specific modifications (Helfrich & Piel, 2016). Engineering approaches targeting acyltransferase domains have further demonstrated the potential to manipulate these pathways for novel compound production (Musiol-Kroll & Wohlleben, 2018). Hybrid NRPS–PKS systems add another layer of complexity, as seen in the biosynthesis of microcystins, which integrate peptide and polyketide elements (Tillett et al., 2000). Insights into these systems have also informed broader efforts to repurpose biosynthetic machinery for industrial applications (Yuzawa et al., 2016).
Despite these advances, a major challenge remains: a large proportion of marine microorganisms are not readily culturable under laboratory conditions. This limitation has historically constrained the discovery of natural products. However, the emergence of genome mining and metagenomics has begun to address this issue, allowing researchers to identify biosynthetic gene clusters (BGCs) directly from environmental DNA (Nikolouli & Mossialos, 2016; Piel et al., 2004). Tools such as antiSMASH and NaPDoS have become central to these efforts, enabling the prediction and classification of gene clusters with increasing accuracy (Blin et al., 2023; Ziemert et al., 2012).
Metagenomic and meta-omic studies have also revealed that many bioactive compounds are produced within complex microbial consortia rather than isolated organisms. The production of ET-743, for instance, has been linked to symbiotic microbial communities associated with marine invertebrates (Schofield et al., 2015; Rath et al., 2011). Similarly, the bry gene cluster identified in uncultured symbionts highlights the importance of ecological context in biosynthesis (Hildebrand et al., 2004; Davidson et al., 2001).
Complementary approaches, including metabolomics and transcriptomics, provide additional layers of insight. Metabolomic profiling of fungi and bacteria has revealed dynamic changes in secondary metabolite production in response to environmental stimuli (Frisvad et al., 2009; Alves et al., 2025). Genome-based analyses have further expanded our understanding of PKS diversity and regulation, particularly in plant-associated and pathogenic fungi (Sayari et al., 2022).
More recently, synthetic biology has begun to reshape the field. Techniques such as CRISPR interference (CRISPRi) enable targeted regulation of biosynthetic pathways, offering new opportunities for activating silent gene clusters and optimizing metabolite production (Choi & Woo, 2020). These developments suggest that the limitations of traditional cultivation-based approaches may gradually be overcome, although challenges related to expression systems and pathway regulation remain.
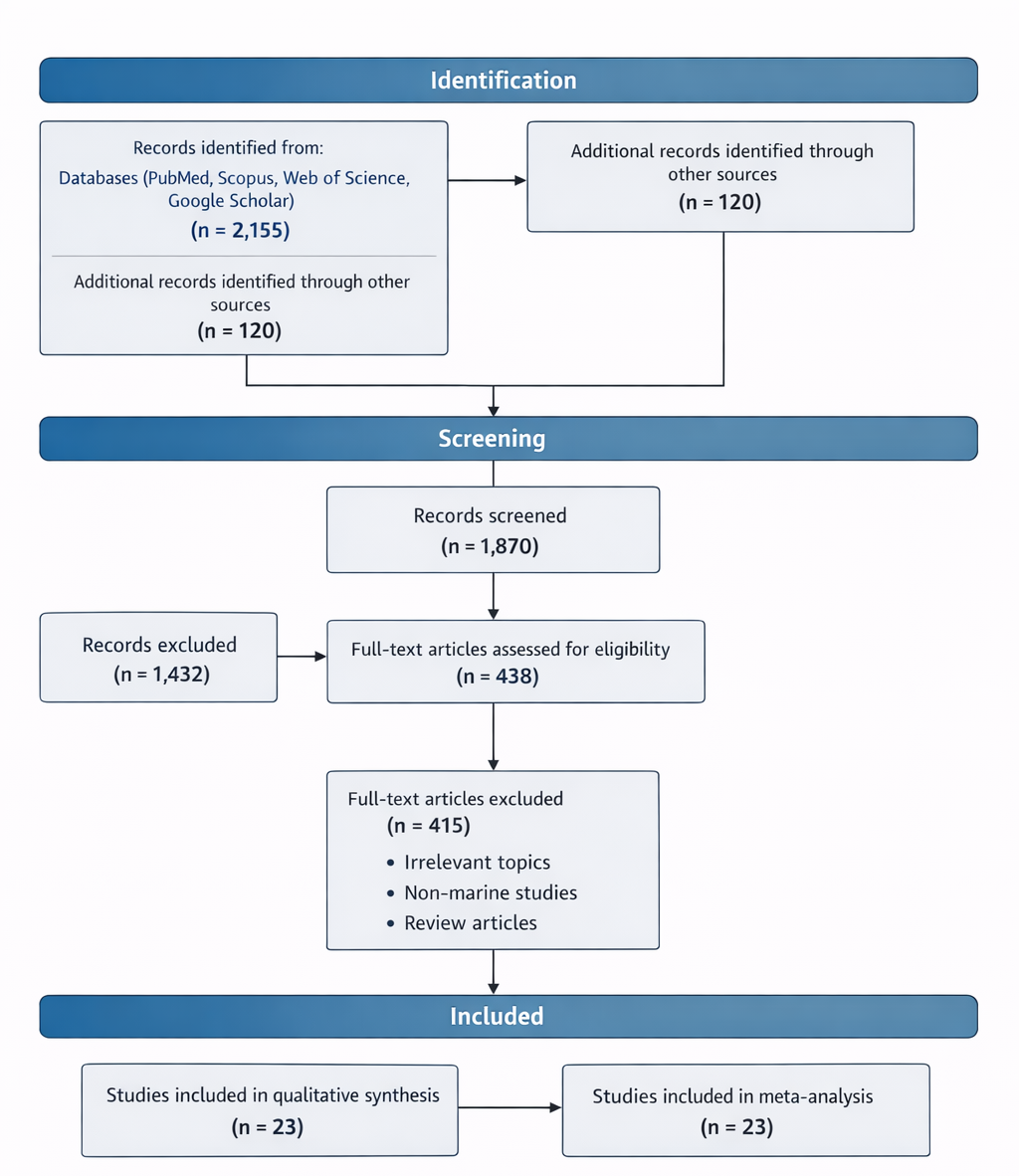
Taken together, these observations point toward a field that is both rapidly evolving and inherently complex. Marine microorganisms represent a vast and largely untapped source of bioactive compounds, yet their study requires the integration of diverse methodologies—from structural biology to bioinformatics and synthetic biology. The present study, therefore, aims to provide a systematic synthesis of existing knowledge, with a particular focus on the diversity, biosynthesis, and bioactivity of marine-derived secondary metabolites. By combining qualitative and quantitative approaches, this review seeks to clarify current trends while also identifying areas where further investigation may be warranted.