1. Introduction
Marine environments—perhaps more than any other biosphere—invite a certain humility when we try to map their biochemical complexity. Beneath fluctuating gradients of light, salinity, temperature, and oxidative stress, microorganisms do not merely survive; they adapt in ways that often feel, at first glance, unexpectedly inventive. Among the metabolic strategies that have drawn sustained attention, carotenoid biosynthesis stands out—not only because of its ecological significance, but also because of its growing relevance to biotechnology and human health (Britton et al., 2004). What is becoming increasingly clear, though not always in a straightforward way, is that marine bacteria contribute a disproportionately rich and structurally diverse subset of these pigments, raising questions about how such diversity arises and how it might be harnessed (Yokoyama et al., 1994; 1995).
Early explorations into bacterial carotenoids, particularly in model organisms such as *Rhodobacter capsulatus*, helped establish a foundational understanding of carotenoid gene clusters and enzymatic steps (Armstrong et al., 1990). These studies, alongside parallel biochemical work dissecting enzymatic transformations (Hundle et al., 1991; Math et al., 1992), defined what is now often referred to as the “base pathway”—a conserved sequence of reactions converting isoprenoid precursors into ß-carotene. The identification and functional characterization of genes such as *crtE*, *crtB*, *crtI*, and *crtY* provided a modular framework that could later be transferred, recombined, and engineered across heterologous systems (Misawa et al., 1990; Sandmann & Misawa, 1992). Yet, even at this early stage, there were hints that bacterial carotenoid biosynthesis might extend beyond the canonical pathways described in plants.
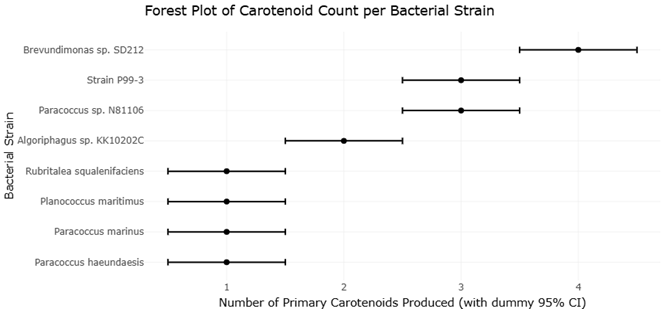
Indeed, when carotenoid pathways were reconstituted in heterologous hosts such as *Escherichia coli*, the system proved both robust and surprisingly flexible (Fraser et al., 1992). Subsequent enzymatic studies refined our understanding of desaturation and cyclization reactions, emphasizing how subtle structural modifications could dramatically alter pigment properties (Schnurr et al., 1996). Functional assignments of individual enzymes further clarified the biochemical logic underpinning carotenoid assembly (Hundle et al., 1992). Still, these insights largely emerged from terrestrial or laboratory-adapted systems, leaving marine microbial diversity comparatively underexplored. It is in marine environments, however, that carotenoid diversity begins to diverge more noticeably from these established norms. Marine bacteria have been shown to produce not only the expected C40 carotenoids, but also structurally unusual compounds—such as C30 diapocarotenoids—that suggest alternative biosynthetic routes (Shindo, Asagi, et al., 2007; Shindo, Endo, et al., 2008). These findings complicate the earlier assumption that carotenoid biosynthesis follows a relatively constrained evolutionary trajectory. Instead, they point toward a more dynamic landscape, where enzymatic innovation appears to be driven, at least in part, by environmental pressures unique to marine ecosystems. The phylogenetic distribution of carotenoid-producing bacteria reinforces this idea. Genera such as *Paracoccus* and *Brevundimonas*, often associated with marine or saline habitats, repeatedly emerge as key producers of structurally complex xanthophylls (Lee et al., 2004; Khan et al., 2008). Comparative taxonomic analyses suggest that these organisms are not isolated curiosities but part of broader evolutionary lineages shaped by both ecological adaptation and, plausibly, horizontal gene transfer (Harker et al., 1998). This raises an interesting possibility: that marine carotenoid pathways may represent optimized biochemical solutions, refined under selective pressures distinct from those encountered in terrestrial systems.
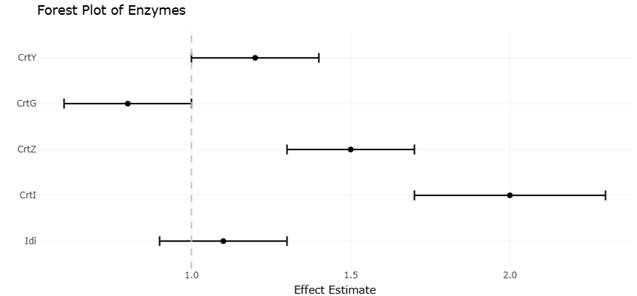
Among the most compelling examples of such optimization is the biosynthesis of ketocarotenoids, particularly astaxanthin (Maruyama, 2007). While ß-carotene itself is widespread, the enzymatic conversion to more functionally elaborate molecules requires additional catalytic steps—most notably ketolation and hydroxylation. These transformations are mediated by enzymes such as CrtW and CrtZ, whose functional expression in bacterial systems has been well documented (Misawa, Yamano, & Linden, 1995a; Misawa, Yamano, & Linden, 1995b). What is striking, however, is the apparent substrate flexibility of these enzymes, which allows them to act on multiple intermediates and generate a spectrum of related compounds (Neudert et al., 1998; Yokoyama et al., 1996a; 1996b). In marine bacteria, this flexibility seems particularly pronounced, suggesting an expanded catalytic repertoire.
The discovery of additional tailoring enzymes further complicates—and enriches—this picture. For instance, the identification of CrtG, a ß-ring 2-hydroxylase, revealed that hydroxylation can occur at positions previously considered sterically inaccessible (Nishida et al., 2005). This finding not only expanded the known diversity of carotenoid structures but also challenged existing assumptions about enzymatic constraints. Similarly, studies on ketocarotenoid biosynthesis have highlighted the role of iterative modifications in generating highly functionalized pigments (Shindo, Kato, & Misawa, 2007). These enzymes, taken together, suggest that marine carotenoid pathways are less linear than once thought, and perhaps better understood as networks of intersecting reactions.
Glycosylation adds yet another layer of complexity. Enzymes such as CrtX mediate the attachment of sugar moieties to carotenoid backbones, producing glycosylated xanthophylls with altered physicochemical properties (Nakagawa & Misawa, 1991; Tao et al., 2006). These modifications are often associated with enhanced membrane stability and stress tolerance, particularly in saline or oxidative environments. Structural studies of such compounds have revealed a level of chemical diversity that is, frankly, difficult to replicate through synthetic chemistry alone (Takaichi et al., 2006). In this sense, marine bacteria function not only as producers of carotenoids but also as natural chemists, capable of generating molecules with finely tuned biological functions.
The functional implications of these pigments extend beyond microbial ecology. A growing body of evidence suggests that carotenoids—especially ketocarotenoids—play roles in human health, including antioxidant protection, cardiovascular support, and potential cancer chemoprevention (Nishino et al., 2002; Jackson et al., 2008). Astaxanthin, in particular, has attracted considerable attention due to its ability to mitigate oxidative damage at the cellular level (Pashkow et al., 2008; Camera et al., 2009). While these effects are often discussed in the context of dietary supplementation, their biosynthetic origins in marine bacteria remain an important, if sometimes underappreciated, aspect of their biological significance.
From an engineering perspective, the modularity of carotenoid pathways has proven especially advantageous. The transfer of marine-derived genes into heterologous hosts has enabled the reconstruction of complete biosynthetic pathways, often with impressive yields (Misawa et al., 1994). These efforts have been further refined through pathway optimization strategies, including the enhancement of precursor supply and the balancing of metabolic flux (Misawa, 2009). Interestingly, some studies suggest that marine enzymes may outperform their terrestrial counterparts in engineered systems, although the reasons for this are not yet fully understood (Teramoto et al., 2003).
Taken together, these observations suggest that marine bacterial carotenoid pathways occupy a unique position at the intersection of ecology, evolution, and biotechnology. They are, in a sense, both conserved and innovative—built upon a shared biochemical foundation, yet capable of generating remarkable structural diversity through relatively subtle enzymatic modifications. This duality makes them particularly attractive targets for systematic investigation.
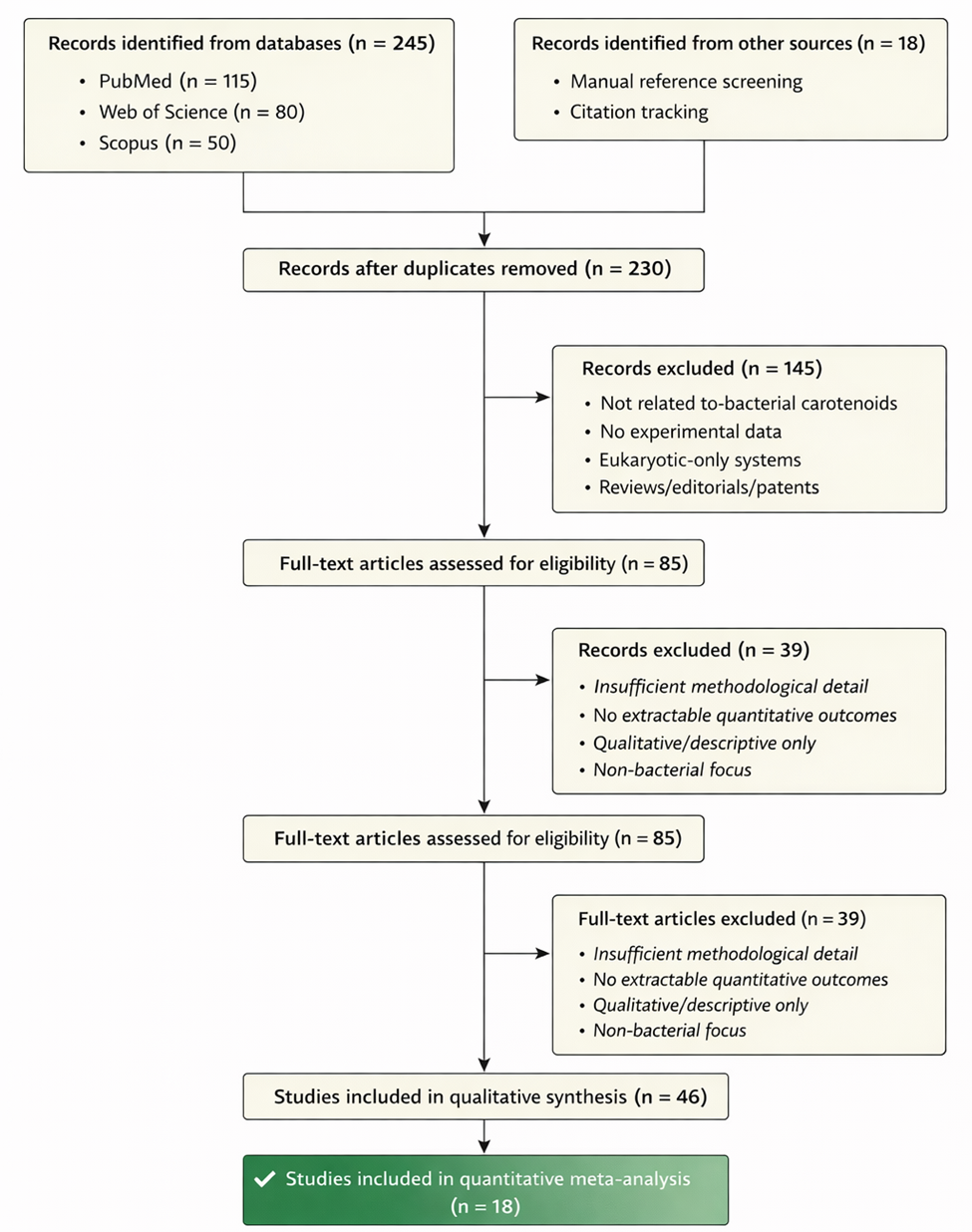
In this context, the present systematic review and meta-analytical synthesis seeks to bring a degree of coherence to what is, admittedly, a complex and sometimes fragmented field. By integrating insights from enzymology, phylogenetics, and metabolic engineering, we aim to clarify how marine bacteria contribute to functional xanthophyll biosynthesis. At the same time, we attempt to identify patterns—whether in enzyme function, pathway organization, or evolutionary distribution—that might inform future efforts to engineer carotenoid production.
Ultimately, understanding marine carotenoid biosynthesis is not merely an academic exercise. It offers a pathway—perhaps not entirely straightforward, but certainly promising—toward the sustainable production of high-value compounds. And in doing so, it underscores a broader point: that even within well-studied metabolic systems, the marine environment continues to reveal layers of complexity that challenge, refine, and occasionally reshape our assumptions.