1. Introduction
The drive for chemical innovation now centers on the rapid discovery of practical, sustainable molecules ready for large-scale production. The depletion of fossil-based resources and the environmental burden of petrochemical industries have intensified efforts to transition toward biologically derived production systems (Sánchez, 2009; Singhvi & Gokhale, 2019). At the same time, medicine faces mounting pressure from antimicrobial resistance and therapy-refractory diseases, underscoring the need for structurally novel bioactive compounds (Aldholmi et al., 2019). These converging pressures have prompted a strategic return to biological systems—not as a retreat from chemistry, but as a recognition that evolutionary processes have generated molecular diversity far beyond conventional synthetic libraries.
Marine microbiomes have increasingly emerged as one of the most promising, yet still insufficiently explored, reservoirs of such diversity (Amoutzias et al., 2016). Oceans represent the largest biome on Earth, encompassing extreme physicochemical gradients—salinity shifts, temperature fluctuations, nutrient scarcity, ultraviolet radiation, and intense ecological competition. Microorganisms thriving in these dynamic conditions have evolved intricate chemical communication and defense systems, often mediated through secondary metabolites (Calteau et al., 2014). Unlike primary metabolites, which are essential for cellular maintenance, secondary metabolites frequently confer ecological advantages—mediating competition, deterring predators, or enabling symbiosis. Many have translated into clinically valuable compounds (Aldholmi et al., 2019; Manavalan et al., 2015).
At the genomic level, these molecules are typically encoded within biosynthetic gene clusters (BGCs). Among the most prominent biosynthetic systems are nonribosomal peptide synthetases (NRPSs) and type I polyketide synthases (PKSs), large modular enzyme complexes that function as molecular assembly lines (Amoutzias et al., 2008; Fischbach et al., 2008). Their domain-based architecture allows combinatorial rearrangement, substrate variation, and evolutionary expansion, thereby generating remarkable chemical plasticity. Comparative genomic analyses reveal that such biosynthetic systems are widely distributed across diverse marine taxa, often with unexpected structural diversity (Cimermancic et al., 2014).
Historically, discovery efforts depended largely on cultivation-based screening. Microorganisms were isolated, cultured, and tested for bioactivity. Although this approach yielded foundational antibiotics and anticancer agents, it captures only a fraction of microbial biosynthetic potential. Many marine microbes resist laboratory cultivation, and even cultivable strains often harbor transcriptionally silent BGCs that remain inactive under standard conditions (Alam et al., 2021). Consequently, traditional methods underestimate true metabolic capacity.
The advent of next-generation sequencing and genome mining reshaped this paradigm. Bioinformatics tools now enable the identification of conserved NRPS and PKS domains directly from genomic sequences, bypassing the need for metabolite detection (Boddy, 2013). Global analyses of prokaryotic genomes have revealed a vast landscape of orphan BGCs with no known products, suggesting an immense reservoir of cryptic chemistry (Cimermancic et al., 2014). Even well-characterized genera frequently possess unexpressed pathways whose functional outputs remain unknown.
Metagenomics further extended this reach by enabling the extraction and sequencing of environmental DNA from complex marine communities (Garza & Dutilh, 2015). This approach has been instrumental in linking several important marine natural products to their microbial symbionts rather than their invertebrate hosts. For example, bryostatin biosynthesis was traced to a bacterial symbiont associated with marine bryozoans (Davidson et al., 2001). Such findings underscore the central role of microbiomes in marine chemical ecology.
Yet, despite technological advances, a persistent challenge remains: the “silent metabolome.” The presence of a biosynthetic gene cluster does not guarantee metabolite production. Environmental cues, regulatory circuits, and ecological interactions strongly influence gene expression. Similar regulatory complexity is well documented in fungal systems, where lignocellulose-degrading enzymes are tightly controlled and environmentally responsive (Hatakka & Hammel, 2010; Kersten & Cullen, 2007). Oxidative enzymes such as fungal peroxygenases and aryl-alcohol oxidases further illustrate how secondary metabolism depends on coordinated extracellular and intracellular regulatory systems (Hernández-Ortega et al., 2012; Hofrichter et al., 2022).
Efforts to activate silent clusters have drawn inspiration from microbial ecology and fungal biotechnology. Approaches such as altering culture parameters (OSMAC strategies), co-cultivation, and targeted genetic manipulation aim to simulate environmental triggers. The importance of extracellular enzyme systems and secreted factors in shaping metabolic output has been demonstrated in fungal interaction studies, where volatile production and enzyme secretion vary with ecological conditions (O’Leary et al., 2019). Secretome regulation studies further highlight how complex transcriptional control governs enzyme deployment (McCotter et al., 2016).
The broader literature on lignocellulose degradation offers instructive parallels. Fungi capable of degrading plant cell walls employ coordinated arrays of carbohydrate-active enzymes and oxidative systems (Blanchette, 1995; Kubicek et al., 2014). Genome-wide analyses of CAZyme repertoires reveal extensive diversification linked to ecological adaptation (Ospina-Giraldo et al., 2010). Secretome predictions in plant-pathogenic fungi demonstrate how lifestyle specialization shapes enzymatic output (Morais do Amaral et al., 2012; Jia et al., 2023). These systems underscore an important point: genomic potential and expressed phenotype are not synonymous. Regulatory context matters.
Similarly, enzymatic systems such as DyP-type peroxidases and laccases exemplify how oxidative metabolism expands functional diversity in fungi (Liers et al., 2010; Loi et al., 2021). These enzymes operate in synergistic networks, illustrating how metabolic capacity emerges from coordinated pathways rather than isolated genes. Comparable principles likely govern NRPS and PKS cluster activation in marine microbes.
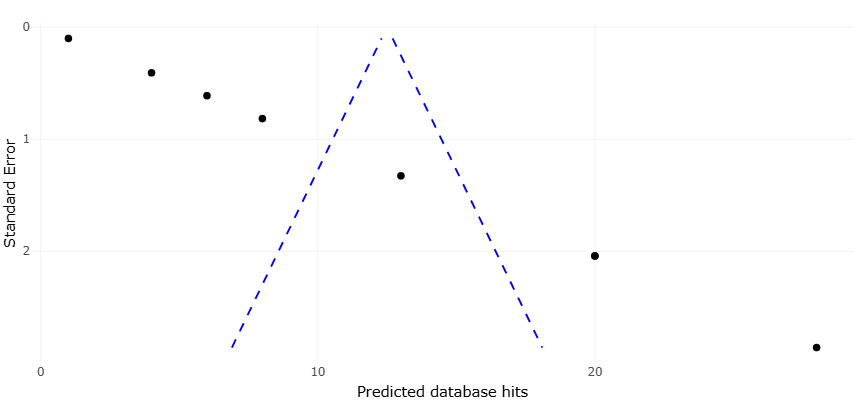
Given this complexity, a key question emerges: how reliably do in silico predictions of biosynthetic capacity translate into experimentally validated metabolite discovery? While some genome mining studies report promising hit rates, others encounter high proportions of inactive clusters (Alam et al., 2021). Variability in sequencing depth, annotation pipelines, cultivation conditions, and assay sensitivity complicates interpretation. Without systematic synthesis, individual findings risk overgeneralization.
A systematic review and meta-analysis provide a structured means to address these uncertainties. By integrating data across genome mining studies, metagenomic surveys, cultivation experiments, and biochemical assays, it becomes possible to evaluate patterns, quantify heterogeneity, and identify reproducible predictors of success. Such synthesis also enables comparison across ecological contexts and taxonomic groups.
This study therefore critically evaluates discovery strategies targeting NRPS- and PKS-derived metabolites within marine microbiomes. Specifically, it assesses (i) the relationship between predicted BGC abundance and observed metabolite or enzymatic activity, (ii) the influence of cultivation and activation conditions on discovery outcomes, and (iii) the effectiveness of genome mining and metagenomic approaches in uncovering novel compounds. By synthesizing genomic, biochemical, and ecological evidence, this work aims to clarify not merely the scale of marine biosynthetic potential, but the practical pathways through which it may be unlocked.