1. Introduction
The natural world is teeming with life forms that remain largely invisible to the naked eye, yet their diversity and biochemical potential are staggering. Microorganisms such as cyanobacteria, microalgae, and uncultured bacteria constitute a significant proportion of global biodiversity, inhabiting ecosystems ranging from sunlit freshwater lakes to the extreme pressures of deep-sea sediments. Historically, our understanding of these microscopic organisms has been constrained by the so-called “great plate count anomaly,” wherein traditional culture-based techniques successfully recover less than 1% of microbial diversity (Handelsman, 2004). Consequently, a substantial fraction of microbial life—commonly referred to as “microbial dark matter”—has remained largely unexplored, leaving untapped reservoirs of novel metabolites and functional potential. Today, a combination of technological advances in sampling, imaging, high-throughput screening, and omics-based analyses has catalyzed a transformative era in microbial exploration, allowing us to systematically uncover, analyze, and exploit this hidden diversity (Alam et al., 2021; Nayfach et al., 2019).
The exploration of extreme and remote habitats has particularly benefited from robotic and autonomous technologies. Deep-sea sediments, hydrothermal vents, and acidic volcanic craters present formidable logistical and environmental challenges, yet they host unique microbial communities with distinctive biochemical capabilities (Gusmão et al., 2023; Crognale et al., 2018). Remotely Operated Vehicles (ROVs) and Autonomous Surface Vehicles (ASVs) have enabled researchers to reach these inaccessible regions, collect samples in situ, and monitor environmental conditions in real time. For instance, ROVs equipped with robotic arms have facilitated the sampling of asphalt ecosystems at depths exceeding 3,000 meters, revealing indigenous cyanobacterial populations in locales previously presumed barren (Gusmão et al., 2023). Likewise, pressure-retaining samplers preserve the physiological state of deep-sea microbes, maintaining in situ conditions for specimens collected at depths of up to 6,000 meters, which is essential for accurate downstream functional studies (Garel et al., 2019).
In parallel, harmful algal blooms (HABs) represent another domain where autonomous monitoring has revolutionized ecological surveillance. These blooms, often driven by nutrient enrichment and climate change, can have devastating effects on aquatic ecosystems, fisheries, and human health. Multi-modal lake and coastal sampling, employing ASVs and Environmental Sample Processors (ESPs), allows near-real-time detection of toxin-producing species such as Pseudonitzschia, coupled with physical and chemical water quality measurements (Salman et al., 2022; Moore et al., 2021). In mesophotic zones off Rapa Nui, filamentous mats have been identified through such approaches, highlighting how robotic surveillance can detect emerging ecological threats that traditional manual sampling might overlook (Sellanes et al., 2021). Furthermore, analogous methodologies have enabled simulations of extraterrestrial exploration, such as Mars drilling missions, using analog environments like the Río Tinto and Solfatara Crater to test the detection of complex microbial biomarkers under extreme acidity (Crognale et al., 2018; Sánchez-García et al., 2020).
Beyond sampling, the characterization of microbial morphology and biochemical features has been transformed by advancements in microscopy. Atomic Force Microscopy (AFM) provides nanometer-resolution three-dimensional imaging of living cells, preserving physiological integrity without chemical fixation (Mišic Radic et al., 2023; Müller & Dufrêne, 2008). This capability allows detailed visualization of cell surfaces, extracellular polymeric substances, and lipid bodies in microalgae such as Parachlorella kessleri, which has implications for sustainable biofuel production (Deniset-Besseau et al., 2021). AFM adaptations, such as Fluidic Force Microscopy (FluidFM), further expand investigative capabilities, enabling the study of interactions between microbubbles and cell surfaces to better understand hydrophobicity and environmental behavior (Demir et al., 2021). Additionally, AFM has elucidated how nanoplastics interact with diatom extracellular matrices, shedding light on environmental pollutant dynamics at the nanoscale (Mišic Radic et al., 2022).
Complementing AFM, cryo-electron microscopy (cryo-EM) has revolutionized high-resolution structural analysis of proteins, photosynthetic complexes, and small molecules (Williamson et al., 2005; Zaharia et al., 2023). By rapidly freezing specimens in vitreous ice, cryo-EM maintains native conformations while avoiding the artifacts introduced by chemical fixation (Yin, 2018; Benjin & Ling, 2020). Notable applications include the elucidation of tetrameric Photosystem I structures in Chroococcidiopsis species and the phycobilisome rod structures in Thermosynechococcus vulcanus, revealing the spatial organization of light-harvesting complexes critical for understanding photosynthetic efficiency (Semchonok et al., 2022; Kawakami et al., 2022). Microcrystal electron diffraction (MicroED) extends these capabilities to small molecules, supporting drug discovery and structural characterization of bioactive compounds produced by marine bacterial symbionts (Danelius et al., 2023; Park et al., 2022).
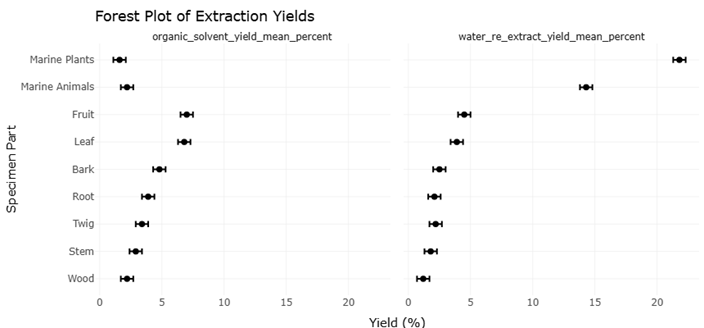
The convergence of microscopy with high-throughput methodologies has enhanced systematic screening of microbial metabolites. High-throughput screening (HTS) platforms, capable of processing up to 100,000 compounds daily, allow rapid evaluation of bioactivity, cytotoxicity, and pharmacological specificity (Martis et al., 2011; Szymanski et al., 2012). High-throughput extraction (HTE) complements these efforts, preserving biologically active molecules from plants, fungi, and marine organisms for subsequent pharmacological evaluation (McCloud, 2010). Libraries such as the National Cancer Institute’s repository, containing over 230,000 extracts, exemplify how large-scale automation accelerates drug discovery pipelines, increasing the likelihood of identifying molecules with antitumor, antimicrobial, or antifungal activity (McCloud, 2010; Wani et al., 1971). Classical examples include Taxol from Taxus brevifolia and jaspamide from marine sponges, whose identification relied on meticulous high-throughput extraction and bioassay-guided fractionation (Wani et al., 1971; Wall et al., 1966).
A paradigm shift has also emerged from culture-independent genomic analyses. Metagenomics allows direct sequencing of environmental DNA, bypassing cultivation barriers and illuminating the metabolic potential of uncultured organisms (Handelsman, 2004; Alam et al., 2021). Shotgun sequencing of marine, soil, and host-associated microbial communities has revealed previously unknown biosynthetic gene clusters (BGCs), linking metabolic traits to their microbial origin (Venter et al., 2004; Owen et al., 2013). Single-cell genomics provides even higher resolution, amplifying the genomes of individual cells through methods such as Multiple Displacement Amplification (MDA), facilitating the functional assignment of metabolites in heterogeneous microbial populations (Stepanauskas, 2012; Blainey, 2013). Such approaches have led to the discovery of unusual polyketide synthase genes from uncultivated marine symbionts and their associated secondary metabolites, which hold promise for novel pharmacological applications (Piel, 2002; Hildebrand et al., 2004).
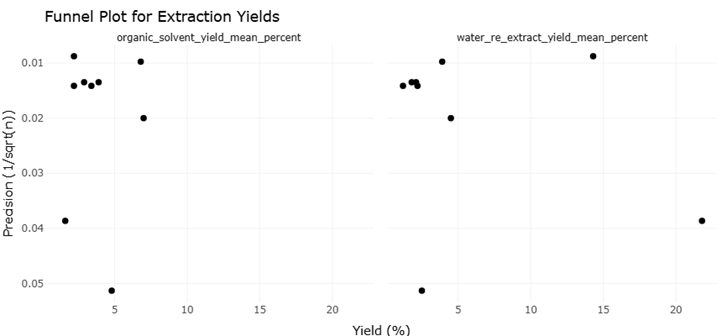
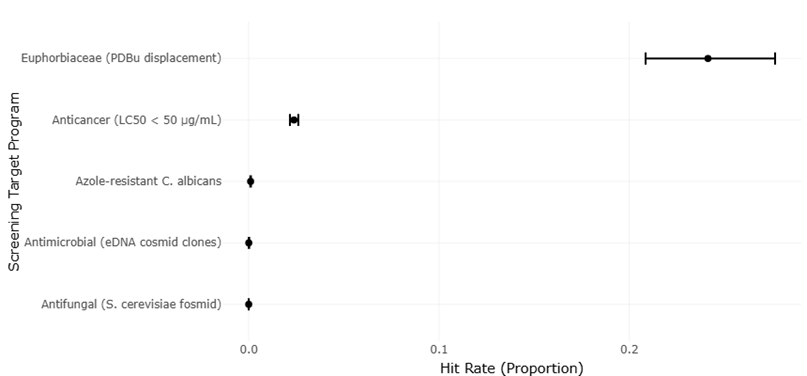
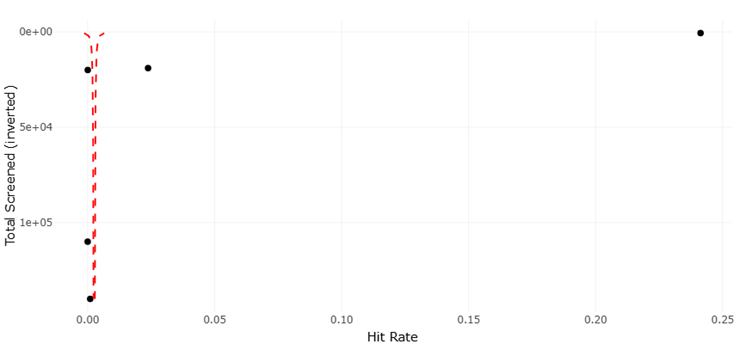
Despite these remarkable advancements, challenges persist. The assignment of biological functions to genes discovered via metagenomics remains complex, particularly for poorly characterized sequences or cryptic biosynthetic pathways (Pessi et al., 2023; Dextro et al., 2021). Variability in sample handling, environmental heterogeneity, and technical limitations in imaging or sequencing can influence results, underscoring the need for systematic evaluation and replication (Singh, 2023; Nayfach et al., 2019). Moreover, while high-throughput systems improve efficiency, the low hit rates observed in large screening programs for antimicrobial and antifungal compounds—sometimes less than 0.01%—highlight the ongoing need for optimized strategies to balance scale with precision (Alley et al., 1988; McCloud, 2010).
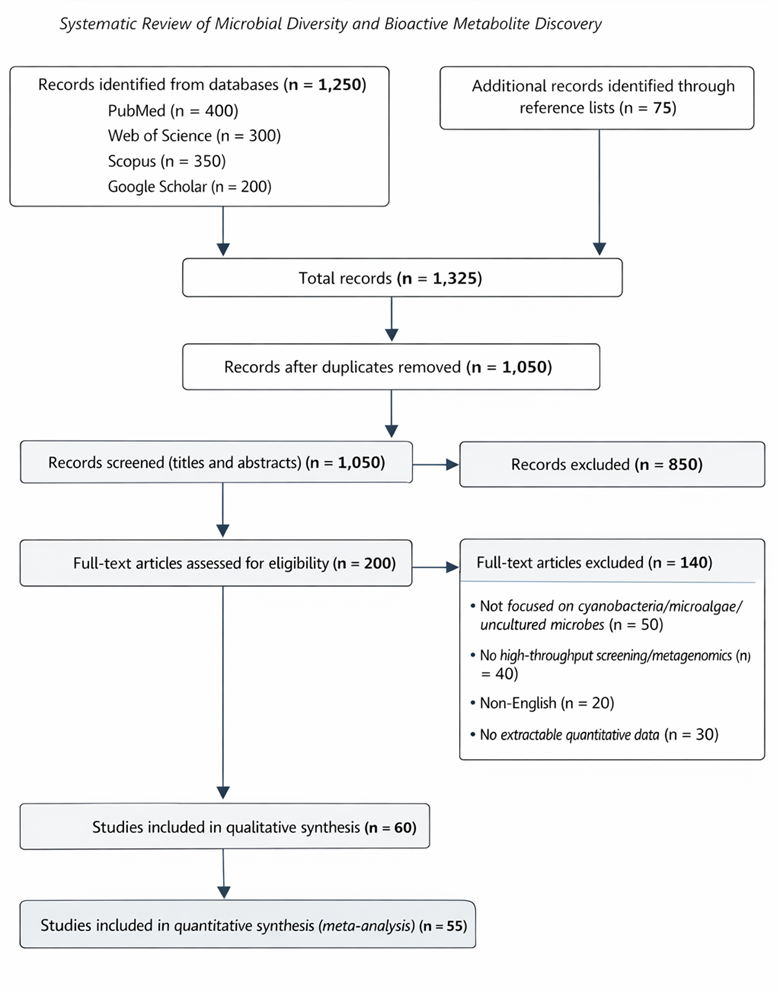
Integrating robotics, microscopy, and omics-based analyses is thus more than a technical exercise; it represents a conceptual shift in our approach to life at the microscale. By combining in situ preservation, single-cell analyses, and high-content functional screening, researchers can now observe microbial communities and their metabolites in their native context, linking ecological roles to biochemical potential (Nishimura et al., 2023; Liu et al., 2022). This holistic framework—bridging sampling, imaging, genomics, and pharmacology—enables systematic reviews and meta-analyses to quantitatively synthesize data across studies, evaluating trends in extraction yields, hit rates, and the discovery of novel bioactive compounds (Martis et al., 2011; Szymanski et al., 2012).
Ultimately, these technological advances transform our perception of microbial life. Where traditional approaches once relied on artificial laboratory cultivation, modern strategies provide a “high-definition window” into the living world, revealing intricate cellular architectures, cryptic metabolic pathways, and ecologically significant interactions as they naturally occur. The combination of robotic sampling, high-resolution imaging, high-throughput bioassays, and multi-omics analysis promises to unlock a new era of natural product discovery, sustainable bioengineering, and a deeper appreciation for the richness of life at the microscale (Zammit et al., 2023; Singh, 2023).