1. Introduction
Ergot alkaloids represent one of the most biologically potent and historically influential classes of fungal secondary metabolites. These nitrogen-containing indole alkaloids are unified by their derivation from L-tryptophan and, with few exceptions, by the presence of a tetracyclic ergoline ring system that underpins their remarkable pharmacological activity (Schiff, 2006; Wallwey & Li, 2011). Interest in ergot alkaloids spans centuries, evolving from fear and superstition to molecular understanding and biotechnological exploitation. Today, these compounds are recognized not only as agents of historical toxicity but also as indispensable tools in modern medicine and valuable models for studying fungal metabolic evolution (Haarmann et al., 2009; Schardl et al., 2006).
The notoriety of ergot alkaloids originates from outbreaks of ergotism, a devastating disease caused by ingestion of cereal grains contaminated with sclerotia of Claviceps purpurea. During medieval Europe, recurring epidemics—collectively referred to as St. Anthony’s Fire—produced symptoms ranging from violent convulsions and hallucinations to ischemic gangrene and limb loss (Schiff, 2006; Haarmann et al., 2009). These effects arise from the ability of ergot alkaloids to mimic endogenous neurotransmitters such as serotonin, dopamine, and adrenaline, enabling them to bind with high affinity to receptors in the nervous system and vasculature (Schardl et al., 2006). While these properties once made ergot a feared contaminant, they later became the foundation for therapeutic innovation.
From a chemical standpoint, ergot alkaloids are traditionally classified into three major structural groups: clavines, lysergic acid amides (ergoamides), and peptide ergot alkaloids (ergopeptines) (Wallwey & Li, 2011; Jakubczyk et al., 2014). Clavines are generally considered the simplest members of this family and often serve as biosynthetic intermediates. Lysergic acid amides represent a further elaboration of the ergoline scaffold, while ergopeptines constitute the most structurally complex group, characterized by a cyclic tripeptide moiety assembled by non-ribosomal peptide synthetases (NRPSs) (Schardl et al., 2006; Gröger & Floss, 1998). This structural diversity directly correlates with biological activity and ecological function, as well as with pharmaceutical utility.
Clinically, ergot alkaloids and their semi-synthetic derivatives continue to play important roles. Methylergometrine is widely used to control postpartum hemorrhage, ergotamine remains a treatment option for acute migraine attacks, and bromocriptine is employed in the management of Parkinson’s disease and hyperprolactinemia (Schiff, 2006; Hulvova et al., 2013). These applications underscore how compounds once associated solely with toxicity have been recontextualized as life-saving medicines. Importantly, such uses have driven intensive research into ergot alkaloid biosynthesis, genetics, and regulation.
Biologically, ergot alkaloids are most strongly associated with fungi in the phylum Ascomycota. While Claviceps species remain the canonical producers, subsequent research has revealed that ergot alkaloid biosynthesis is far more widespread than once believed. Gene clusters responsible for ergot alkaloid synthesis have been identified in endophytic grass symbionts of the genus Epichloë, entomopathogenic fungi such as Metarhizium, and free-living or opportunistic fungi including Aspergillus fumigatus and Penicillium commune (Lorenz et al., 2007; Gao et al., 2011; Panaccione & Coyle, 2005; Kozlovsky et al., 2011). In addition, conserved ergot alkaloid precursor clusters have been detected in the Arthrodermataceae family, suggesting a deep evolutionary origin for this metabolic pathway.
Beyond fungi, ergot alkaloids are also found in higher plants, particularly within the Convolvulaceae and Poaceae families. In these cases, alkaloid presence is typically the result of intimate symbiotic or parasitic relationships with clavicipitaceous fungi rather than autonomous plant biosynthesis (Markert et al., 2008; Ahimsa-Müller et al., 2007). For example, morning glories (Ipomoea spp.) harbor fungal partners that produce ergoline alkaloids, which are vertically transmitted and play defensive roles during early plant development (Beaulieu et al., 2013). These systems highlight ergot alkaloids as ecological currencies that mediate mutualistic interactions and influence host fitness (Schardl et al., 2013).
At the molecular level, ergot alkaloid biosynthesis is one of the most thoroughly characterized fungal secondary metabolic pathways. The pathway begins with the prenylation of L-tryptophan at the C4 position using dimethylallyl diphosphate (DMAPP), a reaction catalyzed by dimethylallyltryptophan synthase (DMATS), encoded by the dmaW gene (Gebler & Poulter, 1992; Tsai et al., 1995). This step commits cellular resources to ergot alkaloid production and establishes the indole-prenyl framework essential for downstream reactions (Floss, 1976; Williams et al., 2000).
Subsequent enzymatic steps lead to the formation of chanoclavine-I aldehyde, a central branch point intermediate conserved across all known ergot alkaloid producers (Jakubczyk et al., 2014; Li & Unsöld, 2006). From this point, pathway divergence occurs, determined by the presence or absence of specific genes within the ergot alkaloid synthesis (EAS) cluster. Species that lack functional NRPS genes terminate the pathway at clavines, whereas those possessing complete NRPS modules proceed to synthesize lysergic acid derivatives and complex peptide alkaloids (Lorenz et al., 2009; Haarmann et al., 2005). Comparative genomic analyses have demonstrated that variation in cluster size and composition reflects both evolutionary history and ecological strategy (Tudzynski et al., 1999; Fleetwood et al., 2007).
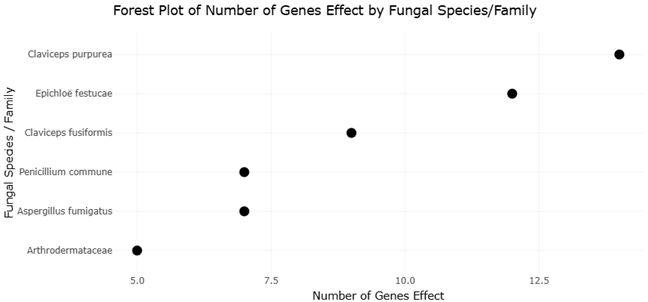
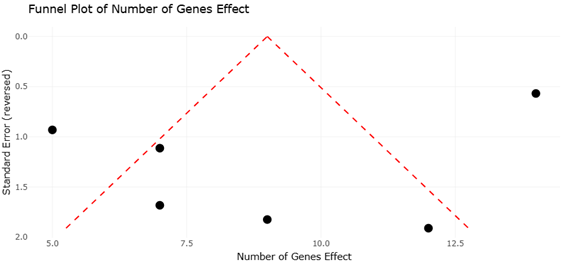
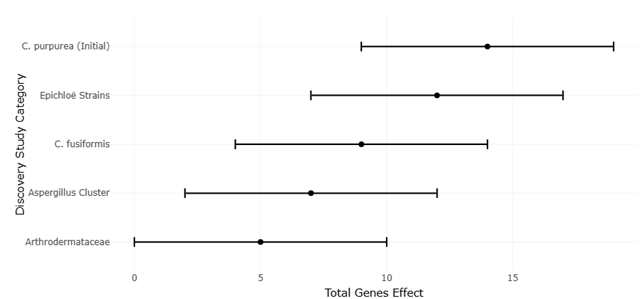
The increasing availability of fungal genome sequences has enabled systematic comparisons of EAS gene clusters across taxa. For instance, Claviceps purpurea harbors a large cluster of approximately 14 genes and produces a diverse spectrum of ergopeptines, whereas Epichloë festucae contains a slightly reduced cluster associated with ergovaline synthesis (Lorenz et al., 2007; Fleetwood et al., 2007). In contrast, Aspergillus fumigatus and Penicillium commune possess smaller clusters that support the production of fumigaclavines, while Arthrodermataceae species appear limited to early pathway intermediates (Coyle & Panaccione, 2005; Kozlovsky et al., 2011). These patterns provide an ideal framework for systematic review and meta-analytical approaches that assess relationships between gene cluster architecture, biosynthetic output, and evolutionary constraint.
From a biotechnological perspective, ergot alkaloid research has entered a new phase driven by genome mining, functional genetics, and enzyme engineering. The overexpression and biochemical characterization of pathway enzymes such as FgaPT2 from A. fumigatus have revealed remarkable substrate flexibility, opening opportunities for chemoenzymatic synthesis of novel prenylated compounds (Unsöld & Li, 2005; Unsöld, 2006). Furthermore, targeted gene deletions and cluster modifications now allow the generation of fungal strains that produce single, pharmaceutically pure alkaloids rather than complex mixtures (Hulvova et al., 2013).
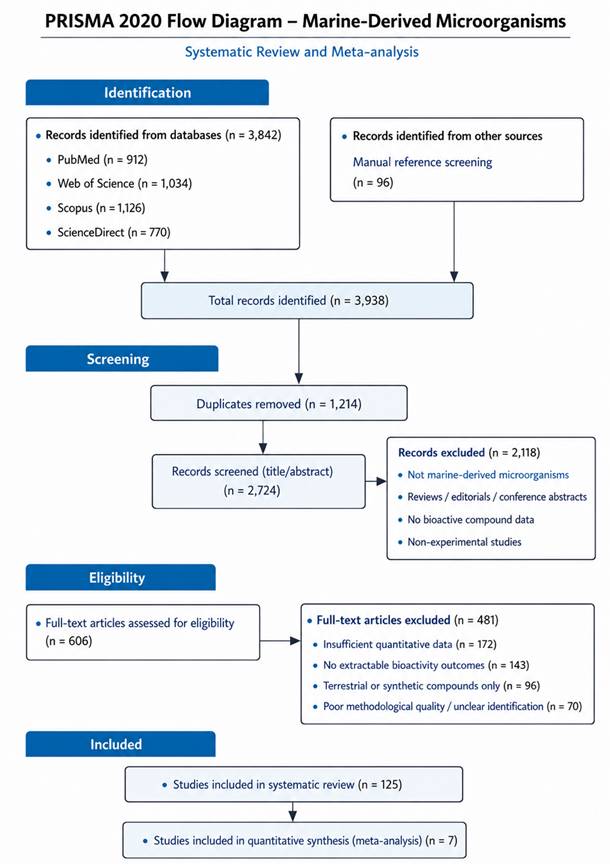
Against this backdrop, the present systematic review and meta-analysis synthesize available molecular, biochemical, and genomic evidence on ergot alkaloid biosynthesis across fungal lineages. By integrating comparative gene cluster data with biosynthetic outcomes, this work aims to clarify evolutionary trends, identify conserved and variable pathway components, and contextualize ergot alkaloids as both ecological mediators and biotechnological resources. In doing so, it reframes ergot alkaloids not merely as relics of historical poisoning, but as dynamic products of fungal metabolism with enduring scientific and medical relevance.