1.Introduction
Cells exist in environments that are rarely static. Nutrient levels fluctuate, oxidative stress rises and falls, and signals from neighboring cells constantly reshape cellular priorities. To cope with this shifting landscape, living systems rely on intricate regulatory frameworks capable of sensing environmental cues and converting them into coordinated biochemical responses. Signal transduction pathways form the core of this adaptive machinery, linking extracellular stimuli with intracellular decision-making processes. Through these networks, cells translate environmental changes into alterations in metabolism, gene expression, and protein function, allowing them to maintain homeostasis while responding to stress or developmental signals (Ruta et al., 2021; Dmitrenko et al., 2022).
At first glance, cellular adaptation might appear to depend largely on transcriptional regulation—simply turning genes on or off as needed. Yet the reality is more nuanced. Transcriptional responses, although powerful, are comparatively slow and energetically demanding. Cells often require mechanisms that operate on much shorter timescales. Post-translational modifications (PTMs) provide such flexibility. By chemically modifying pre-existing proteins, PTMs can rapidly alter protein activity, stability, localization, or interaction networks. In effect, they function as molecular switches that fine-tune cellular behavior without requiring new protein synthesis (Cappadocia & Lima, 2018; Flotho & Melchior, 2013).
Among the many PTMs identified, phosphorylation has historically received the most attention. Protein kinases and phosphatases form complex regulatory circuits that govern almost every aspect of cellular physiology. Advances in phosphoproteomics have revealed just how extensive these regulatory networks are, uncovering thousands of phosphorylation events that collectively shape cellular signaling landscapes (DeMarco & Hall, 2023). In classical signaling cascades—such as the RAS/MAPK pathway—sequential phosphorylation events propagate signals from cell-surface receptors to nuclear transcription factors, enabling cells to respond to growth factors, cytokines, and other extracellular stimuli. Unsurprisingly, disruptions in these pathways frequently contribute to disease, particularly cancer, where aberrant kinase signaling promotes uncontrolled proliferation and survival (Bray et al., 2024; Chen et al., 2023).
However, phosphorylation does not act in isolation. Over the past two decades, it has become increasingly evident that multiple PTMs cooperate to regulate protein behavior. One modification that has attracted considerable interest is SUMOylation, the covalent attachment of small ubiquitin-like modifier (SUMO) proteins to lysine residues on target proteins. Although structurally related to ubiquitination, SUMOylation typically regulates protein interactions, localization, and transcriptional activity rather than directing proteins toward degradation. The SUMO conjugation cycle—mediated by E1-activating, E2-conjugating, and E3-ligating enzymes—creates a tightly controlled system capable of rapidly responding to cellular stress or developmental signals (Geiss-Friedlander & Melchior, 2007; Wild et al., 2024).
SUMOylation has emerged as a central regulatory mechanism in many biological contexts. In nuclear receptor signaling, for example, SUMO modification can alter receptor activity and chromatin interactions, thereby shaping transcriptional outcomes. Dysregulation of SUMO pathways has also been implicated in various cancers, including thyroid malignancies, where altered SUMO enzyme activity contributes to tumor progression and changes in transcriptional regulation (Jiaerken et al., 2024). At the same time, SUMOylation appears to play protective roles in certain physiological contexts. In the nervous system, it participates in maintaining protein homeostasis and may mitigate the formation of toxic protein aggregates associated with neurodegenerative disorders (Anderson et al., 2017).
The functional consequences of PTMs extend beyond protein signaling to encompass broader regulatory processes such as gene expression and RNA metabolism. Signal transduction pathways frequently intersect with RNA processing mechanisms, coordinating transcription with splicing, polyadenylation, and other post-transcriptional events. These interactions allow cells to adjust gene expression programs rapidly and precisely in response to changing conditions (Ruta et al., 2021). Alternative splicing, in particular, plays a crucial role in expanding proteomic diversity. Splicing regulators integrate signals from kinases and chromatin modifiers to determine splice-site selection, thereby generating distinct protein isoforms from the same gene (Black, 2003).
Chromatin-associated regulatory proteins further illustrate the interplay between signaling pathways and transcriptional control. The lysine acetyltransferase p300, for instance, functions as a key transcriptional co-activator that integrates signaling cues with epigenetic regulation. By modifying histones and transcription factors, p300 facilitates enhancer activation and transcriptional initiation. In prostate cancer, interactions between p300 and the androgen receptor promote the expression of oncogenic gene fusions such as TMPRSS2:ERG, highlighting the role of PTM-dependent transcriptional regulation in disease development (Gioukaki et al., 2023).
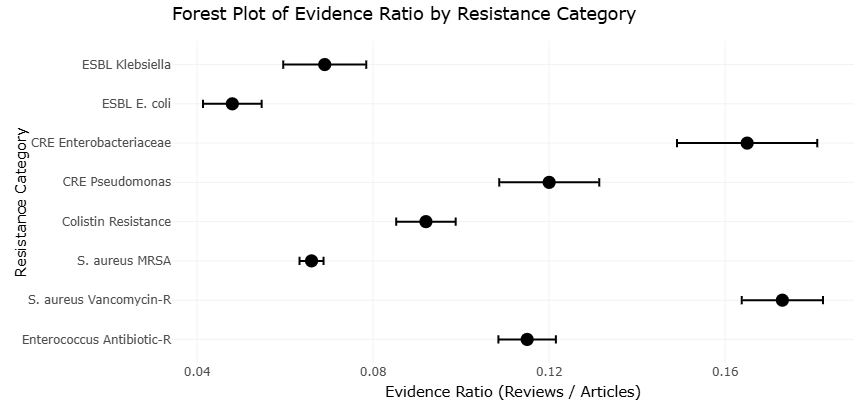
Interestingly, the regulatory logic underlying signal transduction and PTMs is not confined to eukaryotic cells. Microorganisms employ equally sophisticated signaling systems to coordinate their responses to environmental changes. In pathogenic bacteria such as *Staphylococcus aureus*, complex regulatory circuits control virulence, metabolism, and adaptation to host environments. Genome-scale analyses have revealed how these signaling systems contribute to the persistence of multidrug-resistant strains, including methicillin-resistant *S. aureus* (MRSA) (Dmitrenko et al., 2022; Scurtu et al., 2022). The evolutionary success of such pathogens reflects the efficiency with which microbial cells integrate environmental signals into coordinated physiological responses.
Surface proteins and adhesion factors also play critical roles in bacterial signaling and pathogenicity. In *S. aureus*, these molecules mediate interactions with host tissues and immune defenses, facilitating colonization and infection (Foster et al., 2014; Wang & Matunis, 2024). Over time, the accumulation of genetic and regulatory adaptations has led to the emergence of highly successful lineages of MRSA that spread globally in both healthcare and community settings (Enright et al., 2002). Such examples illustrate how signal transduction networks drive microbial evolution and adaptation.
Beyond signaling itself, PTMs have also shaped the discovery of bioactive natural products. Ribosomally synthesized and post-translationally modified peptides (RiPPs) represent a remarkable class of biomolecules whose biological activity often depends on complex PTM-mediated structural modifications. These peptides exhibit diverse pharmacological properties, including antimicrobial and anticancer activities (Arnison et al., 2013; Luo & Dong, 2019). Recent research has highlighted plant-derived RiPPs as promising therapeutic candidates, demonstrating potent cytotoxic effects against tumor cells while offering new opportunities for drug development (Hwang et al., 2025).
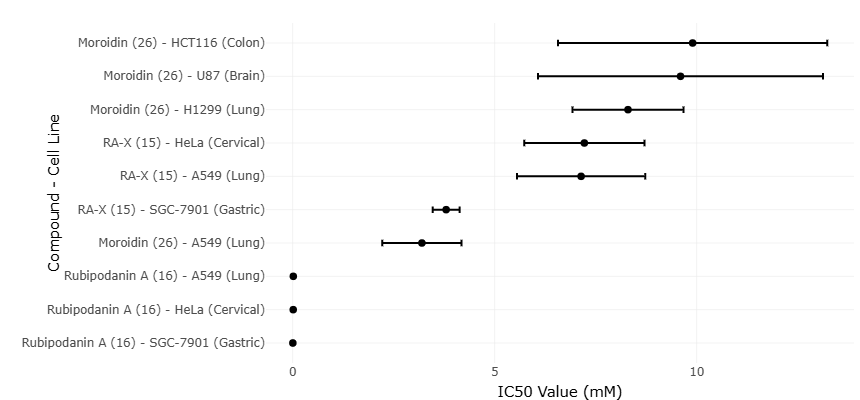
Specific cyclopeptides illustrate how PTM-driven structural modifications can influence biological function. Compounds such as rubipodanin A and moroidin exhibit distinctive conformations and bioactivities that arise from subtle chemical modifications during biosynthesis (Wang et al., 2015; Xu et al., 2022). These molecules provide compelling examples of how nature exploits PTMs not only for regulatory signaling but also for generating chemically diverse metabolites with therapeutic potential.
From an evolutionary perspective, the emergence of PTM-based regulation may have deep biochemical roots. Even simple peptides—once thought to be largely inert—can display catalytic or regulatory properties. Studies of primitive peptide systems suggest that short sequences may have served as early catalysts in prebiotic environments, potentially laying the groundwork for the sophisticated enzymatic networks observed in modern cells (Wieczorek et al., 2017; Sangiorgio et al., 2024). Although speculative, such findings hint that contemporary signaling pathways may reflect ancient biochemical strategies refined over billions of years.
The biomedical significance of these regulatory systems is becoming increasingly clear. Diseases ranging from cancer to neurodegeneration often arise when signaling networks lose their delicate balance. In neurodegenerative disorders, for instance, failures in protein quality control mechanisms allow misfolded proteins to accumulate and aggregate within neurons. PTMs, including SUMOylation and phosphorylation, are deeply involved in maintaining proteostasis and directing damaged proteins toward refolding or degradation pathways (Gestwicki & Garza, 2012).
Taken together, these observations underscore the central role of signal transduction and post-translational modifications in cellular adaptation. Rather than acting as isolated mechanisms, signaling pathways, PTMs, transcriptional regulators, and metabolic networks form an integrated regulatory system that allows cells to respond dynamically to environmental and physiological challenges. In this narrative review, we examine current insights into how these interconnected mechanisms govern cellular adaptation across diverse biological contexts—from microbial pathogenesis to cancer biology and neurodegenerative disease. By synthesizing evidence from molecular biology, biochemistry, and systems biology, we aim to highlight the unifying principles that underlie signaling-based regulation and to identify emerging directions for future research.