1. Introduction
Biomedical signal analysis—once grounded largely in manual interpretation and relatively rigid signal-processing pipelines—has, over the past decade, entered a period of rapid and somewhat uneven transformation. The integration of artificial intelligence (AI) and machine learning (ML) has not simply improved performance metrics; it has begun to reshape how clinical signals are conceptualized, interpreted, and ultimately used in decision-making. Electroencephalography (EEG), electrocardiography (ECG), electromyography (EMG), and respiratory signals, long treated as domain-specific diagnostic tools, are increasingly being reinterpreted as rich, high-dimensional data sources capable of supporting predictive and even anticipatory models of disease (Alqudah & Moussavi, 2025). Yet, despite this momentum, the transition from methodological innovation to reliable clinical utility remains, in many respects, incomplete.
It is tempting to frame AI-driven biomedical signal analysis as a straightforward success story—after all, reported accuracies in controlled settings are often strikingly high. Deep learning architectures, particularly convolutional neural networks and hybrid transformer-based models, have demonstrated strong performance in detecting subtle neurological and psychiatric patterns from EEG data, including early indicators of relapse in complex conditions such as schizophrenia (Yasin et al., 2025). Similarly, cardiology has seen substantial advances, where ML models trained on annotated ECG datasets can achieve near-human or even superhuman classification accuracy in arrhythmia detection. These developments suggest not only technical progress but also a shift in epistemology: from rule-based interpretation toward data-driven inference.
However, a closer reading of the literature introduces a degree of hesitation. High performance in controlled datasets does not necessarily translate into robustness across diverse clinical contexts. In drug discovery, for instance, AI has accelerated candidate screening and molecular prediction pipelines, but questions remain regarding the clinical validity of these outputs and their reproducibility in real-world settings (Dermawan & Alotaiq, 2025; Higgins & Green 2011; Huedo-Medina et al. 2006). A similar pattern emerges in perinatal and obstetric applications, where AI systems show promise in predictive modeling yet often lack comprehensive evaluation across heterogeneous populations (El Arab et al., 2025). These inconsistencies highlight a broader tension: the distinction between algorithmic capability and clinical trustworthiness.
Beyond individual applications, emerging paradigms such as digital twin cognition further complicate this landscape. By integrating biomarker data, computational modeling, and AI-driven inference, digital twin systems aim to simulate patient-specific physiological states. While conceptually compelling, their empirical grounding remains variable, often constrained by limited datasets and methodological heterogeneity (Gkintoni & Halkiopoulos, 2025). In dermatologic AI, for example, diagnostic systems for melanoma have achieved impressive accuracy under controlled conditions, yet persistent concerns about bias—particularly across skin tones—underscore the fragility of these models when exposed to real-world variability (Górecki et al., 2025).
Taken together, these developments suggest that the field is not merely evolving—it is, perhaps, negotiating its own boundaries. The promise of AI in biomedical signal analysis lies not only in improved predictive performance but also in its potential to uncover latent physiological patterns. Yet this promise is tempered by recurring methodological challenges: small sample sizes, lack of external validation, inconsistent preprocessing protocols, and, not infrequently, selective reporting of outcomes (Chan et al., 2004). These issues are not trivial; they directly influence the interpretability, reproducibility, and ultimately the clinical adoption of AI systems.
In this context, systematic review and meta-analysis emerge not as optional tools but as essential methodological frameworks. The synthesis of heterogeneous studies requires careful statistical treatment, particularly when effect sizes vary across designs, populations, and analytical pipelines. Foundational work in meta-analysis has long emphasized the importance of accounting for between-study variability, whether through fixed- or random-effects models (DerSimonian & Laird, 1986; Hedges & Olkin, 1985). More recent perspectives further stress the need to evaluate heterogeneity explicitly, using metrics such as I² and related statistics, to distinguish genuine variability from sampling error (Higgins et al., 2003; Higgins & Thompson, 2002).
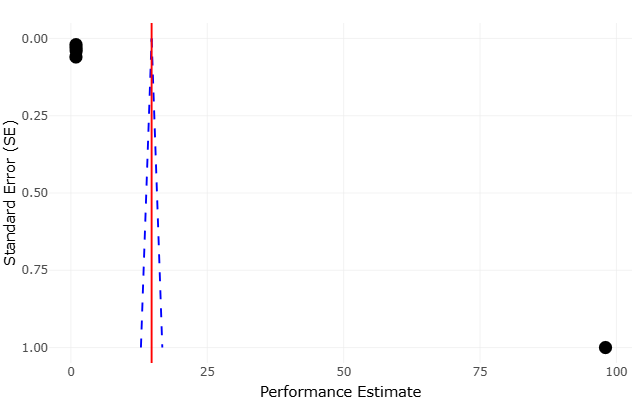
At the same time, meta-analytic rigor extends beyond statistical modeling. Issues such as publication bias, small-study effects, and outcome reporting bias must be carefully assessed to avoid inflated or misleading conclusions (Egger et al., 1997; Harbord et al., 2006). The broader methodological literature reminds us that evidence synthesis is as much about critical appraisal as it is about aggregation (Cooper et al., 2009; Borenstein et al., 2011). Indeed, early reflections on research synthesis emphasized that summarizing evidence requires not only technical precision but also interpretive caution, particularly when integrating findings from diverse and sometimes incompatible study designs (Light & Pillemer, 1984).
These considerations are especially relevant in AI research, where rapid publication cycles and evolving methodologies can obscure underlying limitations. The dependence of effect sizes across studies, for instance, introduces additional complexity, as shared datasets or overlapping methodologies may violate assumptions of independence (Gleser & Olkin, 2009). Similarly, the interpretation of heterogeneity and bias tests requires nuance; statistical significance alone does not necessarily imply meaningful inconsistency or distortion (Ioannidis, 2008).
Moreover, the assessment of evidence quality has gained increasing prominence, particularly in clinical and translational research. Frameworks such as GRADE provide structured approaches to evaluating the strength and certainty of evidence, taking into account study design, consistency, directness, and potential biases (Guyatt et al., 2008). When applied to AI-driven biomedical studies, such frameworks reveal a recurring pattern: strong internal performance paired with limited external validation and uncertain generalizability. This gap, while not unique to AI, is arguably amplified by the complexity and opacity of modern machine learning models.
Another layer of complexity arises from the statistical underpinnings of meta-analysis itself. Classical approaches, including variance-based weighting and aggregation of standardized effect sizes, remain foundational (Fleiss, 1993; Greenland & O’Rourke, 2001). Yet, as datasets grow more complex and interconnected, newer considerations—such as correcting for measurement error or addressing non-independence—become increasingly relevant (Hunter & Schmidt, 2004). The challenge, then, is not merely to apply established methods but to adapt them thoughtfully to the evolving characteristics of AI research.
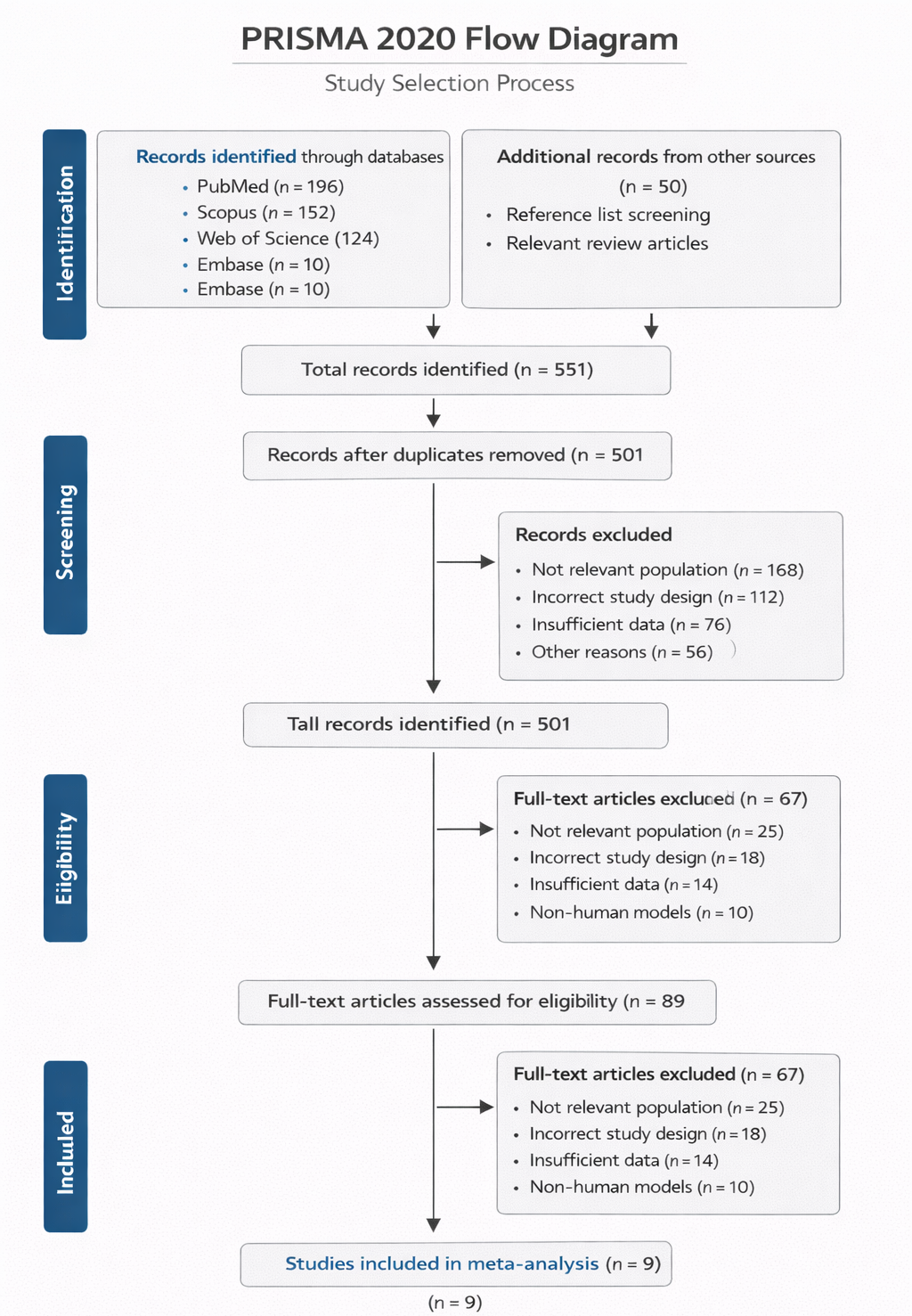
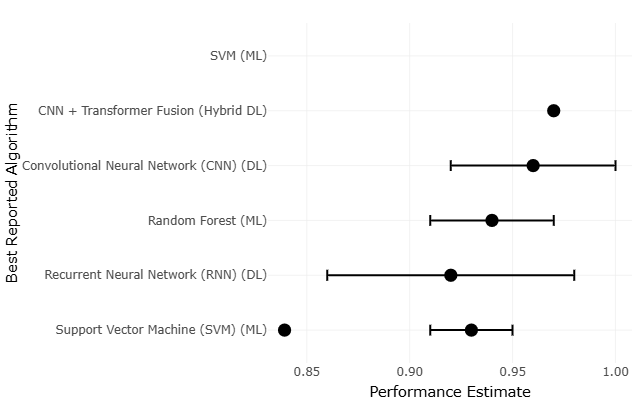
In light of these challenges, the present study adopts a systematic review and meta-analytic approach to examine AI and ML applications in biomedical signal analysis across multiple domains. By synthesizing evidence from EEG, ECG, EMG, and respiratory signal studies, this review seeks to move beyond isolated performance metrics and toward a more integrated understanding of methodological trends, validation practices, and clinical applicability. Quantitative synthesis methods, informed by established principles of research integration, enable the comparison of effect sizes and performance outcomes across heterogeneous studies (Lau et al., 1997). At the same time, qualitative assessment provides context—highlighting where methodological rigor is strong and where it may be, perhaps, less certain.
Ultimately, this work is motivated by a simple but pressing question: how reliable are current AI-driven approaches in biomedical signal analysis when viewed collectively rather than individually? The answer, as this review suggests, is neither wholly optimistic nor dismissive. Instead, it lies somewhere in between—shaped by impressive technical achievements, tempered by methodological limitations, and guided by the ongoing need for transparency, validation, and careful interpretation.