1. Introduction
Liver diseases continue to represent a substantial and persistent burden on global public health. Across both developed and developing regions, disorders affecting the liver contribute significantly to morbidity, long-term disability, and premature mortality. The liver itself is an extraordinary organ—indeed, the largest internal gland in the human body—and its physiological responsibilities are remarkably diverse. It regulates metabolic homeostasis, converts nutrients into usable biochemical forms, detoxifies harmful compounds, synthesizes essential proteins, and supports immune defense mechanisms against invading pathogens (Dritsas & Trigka, 2023). Because these processes are deeply integrated with almost every physiological system, even modest impairment of hepatic function can lead to complex systemic consequences. Viral infections, alcohol consumption, metabolic disorders, environmental toxins, and lifestyle-related risk factors all contribute to the progressive deterioration of liver health, often culminating in conditions such as cirrhosis, fatty liver disease, or hepatocellular carcinoma. Despite the medical importance of early detection, diagnosing liver disease at its initial stages remains difficult. In many cases, symptoms appear only after significant physiological damage has already occurred. Traditional diagnostic approaches—such as imaging techniques, liver biopsies, and biochemical assays—are undeniably valuable, yet they can be costly, time-consuming, or invasive. Moreover, these procedures often identify disease only after measurable structural damage has taken place. This challenge has prompted researchers to explore alternative strategies capable of detecting subtle patterns within clinical data that may indicate early disease risk. In this context, machine learning has gradually emerged as a promising analytical paradigm capable of transforming the landscape of medical diagnostics (Ding et al., 2022; Khan et al., 2022). Machine learning models possess the capacity to analyze complex, high-dimensional datasets and uncover relationships that may not be immediately visible through conventional statistical approaches. Clinical datasets often contain numerous biochemical indicators, demographic variables, and laboratory measurements whose interactions are nonlinear and multifaceted. Human interpretation alone may struggle to identify meaningful relationships among these variables. Machine learning algorithms, however, can evaluate large volumes of data simultaneously, learning from patterns embedded within them and generating predictive models capable of estimating disease probability with considerable accuracy. Over the past decade, such techniques have increasingly been applied to healthcare problems ranging from cancer detection to cardiovascular risk prediction, demonstrating encouraging results across multiple domains (Ding et al., 2022; Khan et al., 2022). Within the specific context of liver disease, several studies have explored the potential of machine learning algorithms for improving diagnostic performance and clinical decision-making. These approaches typically rely on patient clinical profiles—including laboratory markers such as bilirubin levels, liver enzyme concentrations, and demographic characteristics—to train predictive models capable of distinguishing between healthy individuals and those with hepatic disorders. The capacity of machine learning systems to process these diverse inputs simultaneously makes them particularly well suited for this task. In fact, research has shown that certain machine learning algorithms can achieve impressive diagnostic performance when applied to liver disease prediction problems (Mostafa et al., 2021). These developments suggest that data-driven predictive frameworks may serve as valuable decision-support tools in clinical environments where early identification of risk factors is essential.
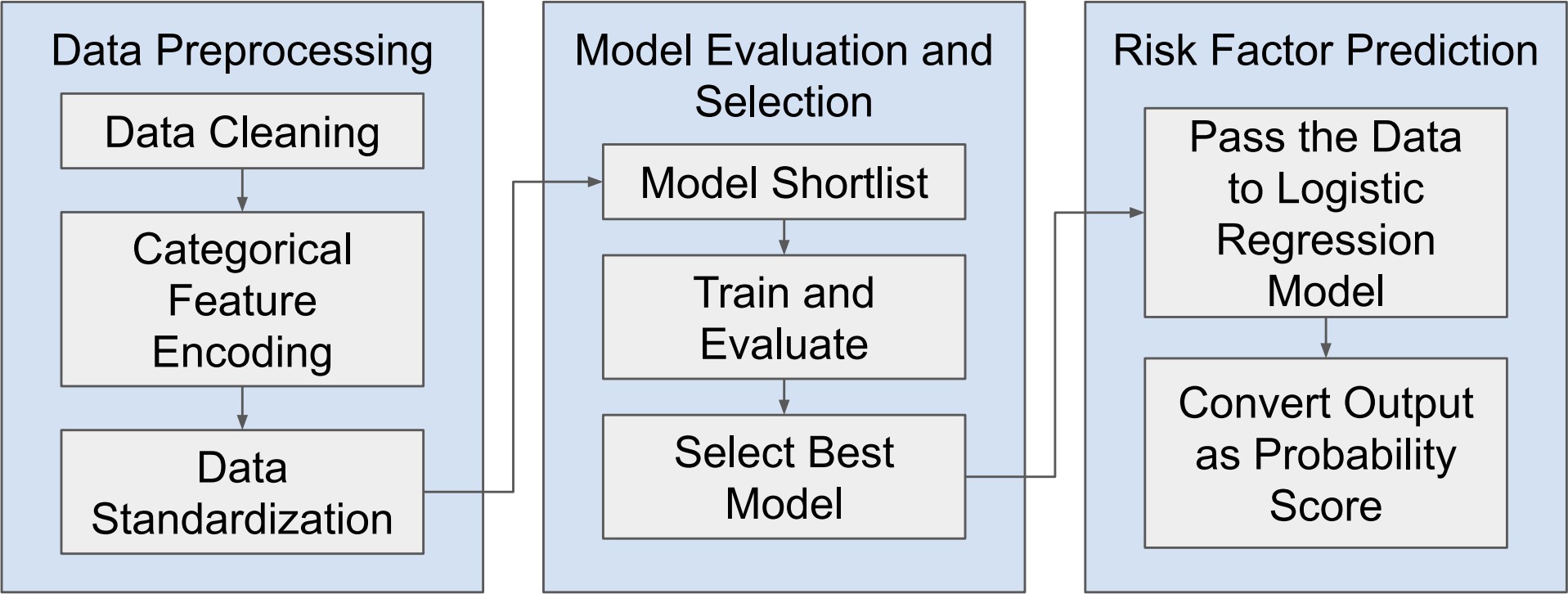
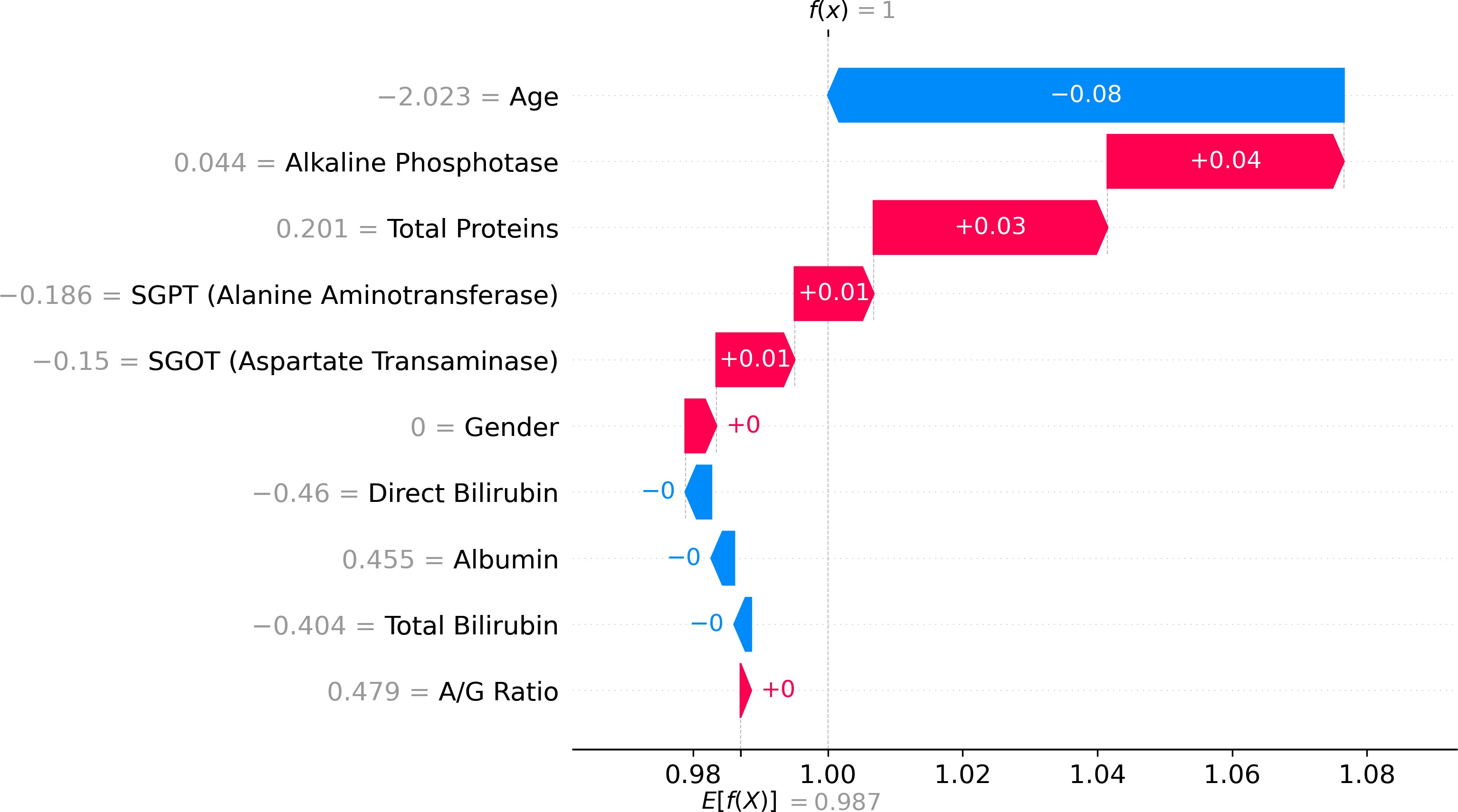
Nevertheless, as machine learning models become increasingly sophisticated, a new challenge has emerged—interpretability. Many advanced predictive models, particularly those based on ensemble methods or deep learning architectures, are often described as “black boxes.” While such models may achieve high predictive accuracy, their internal decision-making processes can be difficult to interpret. In medical contexts, this lack of transparency can raise serious concerns. Healthcare professionals must understand not only what a model predicts but also why it reaches a particular conclusion. Without such understanding, clinicians may hesitate to rely on algorithmic recommendations when making critical diagnostic or therapeutic decisions. This growing concern has led to a parallel research movement focused on explainable artificial intelligence (XAI). Explainability aims to provide insights into how machine learning models generate predictions, thereby improving transparency, trust, and accountability in automated decision systems. Among the various techniques developed to address this issue, SHAP (SHapley Additive exPlanations) has gained particular prominence. Derived from cooperative game theory, SHAP values quantify the contribution of each feature to an individual prediction, allowing researchers and clinicians to interpret model behavior in a consistent and mathematically grounded manner (Stenwig et al., 2022). By decomposing predictions into feature contributions, SHAP makes it possible to identify which variables exert the strongest influence on model outcomes. The importance of explainability in healthcare extends beyond technical transparency. Interpretable models can reveal clinically meaningful insights that might otherwise remain hidden within complex datasets. For example, identifying which biochemical markers contribute most strongly to liver disease prediction can enhance clinical understanding of disease progression and support more targeted diagnostic strategies. Explainable machine learning can therefore serve not only as a predictive tool but also as a mechanism for generating new medical knowledge. From ethical and regulatory perspectives, explainability is equally critical. Healthcare systems must ensure that algorithmic decisions are fair, understandable, and aligned with established clinical principles (Amann et al., 2020). In this study, the identification of hidden risk factors associated with liver disease is investigated using the Indian Liver Patient Dataset (ILPD), a widely used clinical dataset available through the UCI Machine Learning Repository (Ramana & Venkateswarlu, 2012). The ILPD dataset contains multiple biochemical indicators and demographic variables associated with liver health, making it a suitable benchmark for evaluating predictive models. Yet, as with many medical datasets, the relationships between these variables and disease outcomes are complex. Some predictors may exert strong direct effects, whereas others may interact in subtle ways that are not immediately obvious through traditional analytical approaches. To address this challenge, the present research integrates machine learning modeling with explainable artificial intelligence techniques. The dataset undergoes rigorous preprocessing and exploratory analysis before being used to train predictive models capable of distinguishing between liver disease patients and healthy individuals. Subsequently, SHAP-based interpretability methods are applied to analyze model behavior and identify the most influential features contributing to prediction outcomes. Through this combined approach, the study seeks not only to achieve accurate classification performance but also to illuminate the underlying risk factors that may contribute to liver disease development. Understanding these determinants holds important implications for clinical practice. If certain biochemical indicators consistently emerge as dominant predictive features, they may serve as valuable diagnostic markers in routine medical screening. Furthermore, insights derived from interpretable machine learning models can support clinicians in making evidence-based decisions, improving both diagnostic accuracy and patient management strategies. The remainder of this paper is organized as follows. Section II reviews previous research related to machine learning applications in liver disease diagnosis and prediction. Section III presents the dataset description, preprocessing procedures, exploratory data analysis, model development, evaluation methods, and interpretability framework used in this study. Section IV discusses the experimental results and their implications. Finally, Section V concludes the paper and outlines potential directions for future research in explainable machine learning for medical diagnostics.